Gram Stain (Mixed Bacteria)
Gram Stain (Mixed Bacteria)
MicrobiologyHeat-fixed bacterial smear with Gram staining showing gram-positive and gram-negative organisms
Drag & drop images here to add to this specimen
or click Upload Image aboveRelease to upload
Image Library
32 images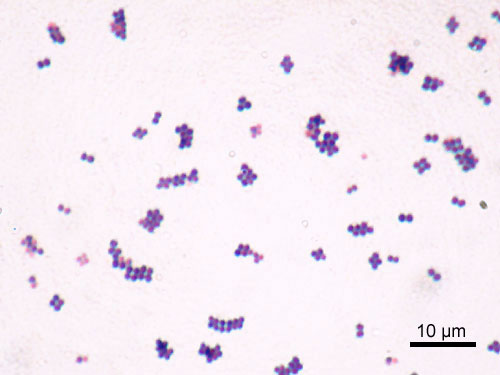
Gram-stained smear of Staphylococcus aureus showing characteristic grape-like clusters of gram-positive cocci (dark purple).
Oil immersion — classic staphylococcal cluster morphology
Wikimedia Commons — CC BY-SA 3.0
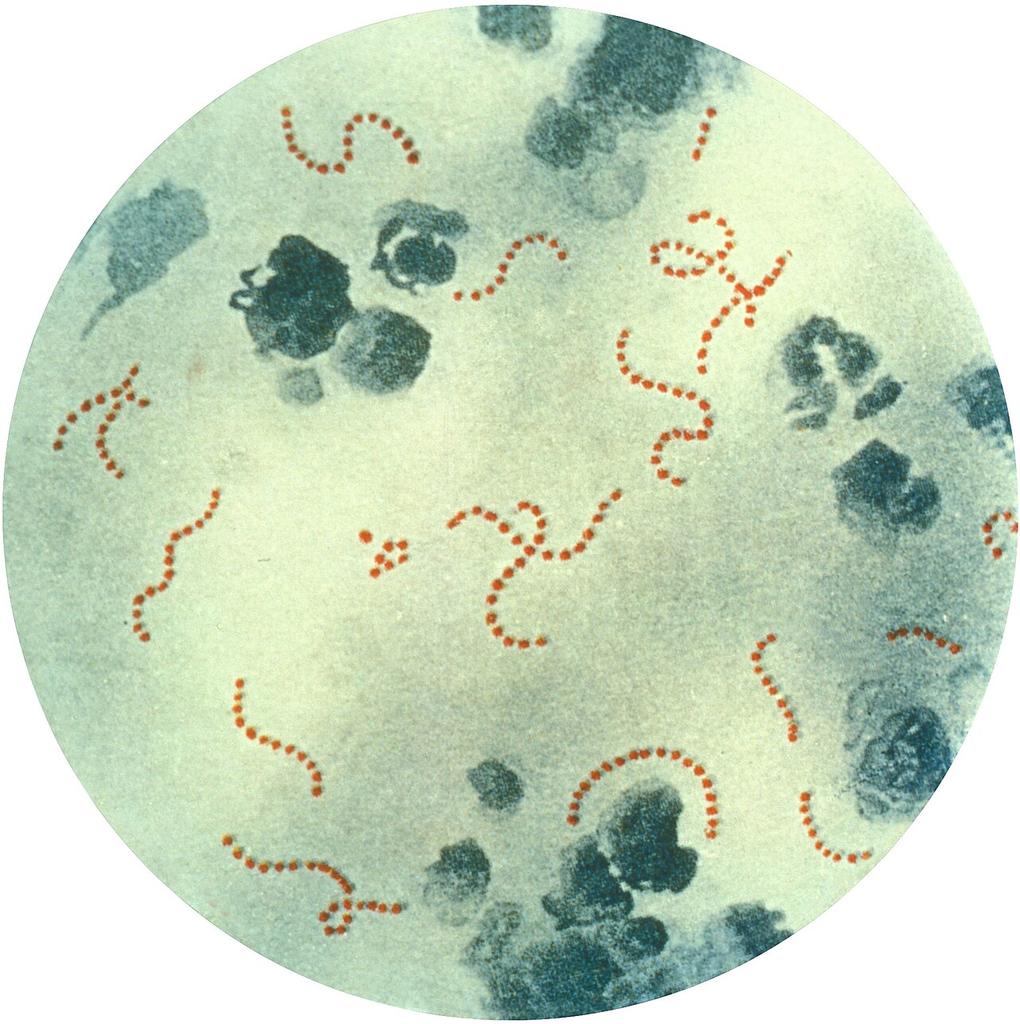
Gram-stained smear of Streptococcus pyogenes (Group A Strep) showing gram-positive cocci in chains.
Oil immersion — chains of cocci, group A beta-hemolytic streptococcus
Wikimedia Commons — CC BY-SA 3.0
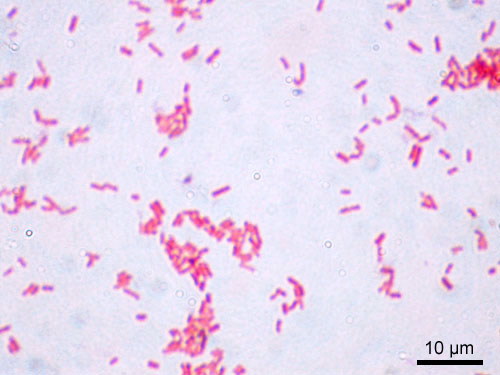
Gram-stained smear of Escherichia coli showing pink/red gram-negative bacilli. Uniform rod-shaped bacteria characteristic of Enterobacteriaceae.
Oil immersion — gram-negative rods, most common clinical isolate
Wikimedia Commons — CC BY-SA 3.0
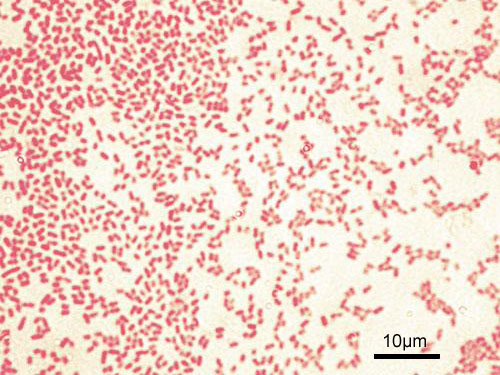
Gram stain showing slender gram-negative bacilli of Pseudomonas aeruginosa. Organisms appear as thin pink rods.
Oil immersion — slender gram-negative rods, opportunistic pathogen
Wikimedia Commons — CC BY-SA 3.0
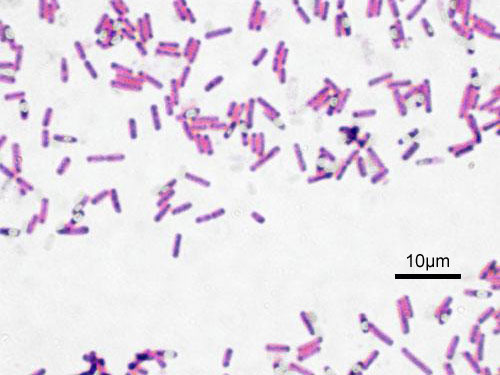
Gram-stained smear of Bacillus subtilis showing large gram-positive bacilli in chains. Spore-forming rod with subterminal endospores sometimes visible.
Oil immersion — large gram-positive rods in chains, spore-forming
Wikimedia Commons — CC BY-SA 3.0
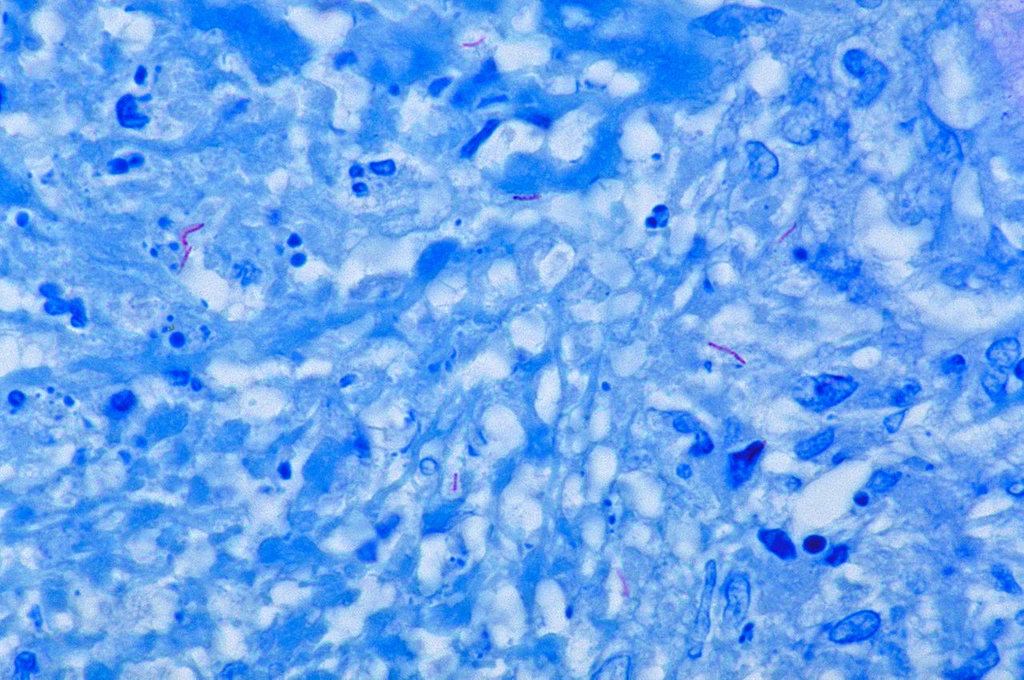
Ziehl-Neelsen acid-fast stain showing bright red acid-fast bacilli (AFB) of Mycobacterium tuberculosis against blue counterstain background.
Oil immersion — acid-fast bacilli, comparison with Gram stain
Wikimedia Commons — Public Domain (CDC)
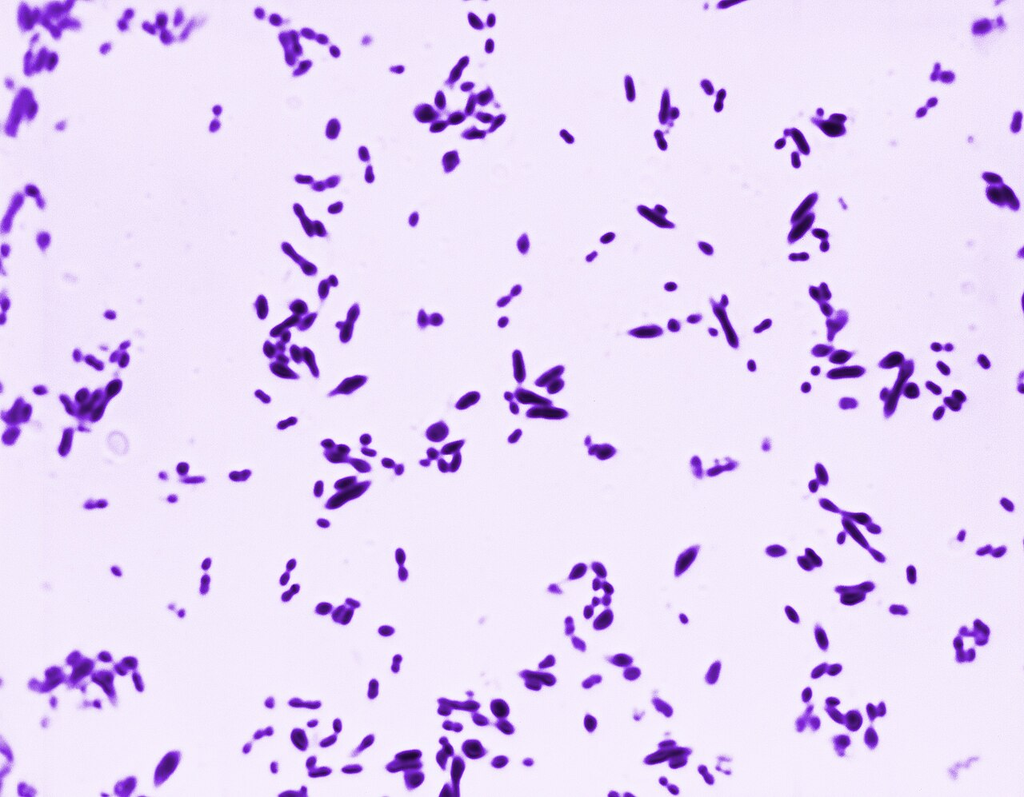
Gram stain of Streptococcus pneumoniae showing lancet-shaped gram-positive diplococci with characteristically pointed ends.
Oil immersion — lancet-shaped diplococci, common cause of pneumonia
Wikimedia Commons — Public Domain (CDC)
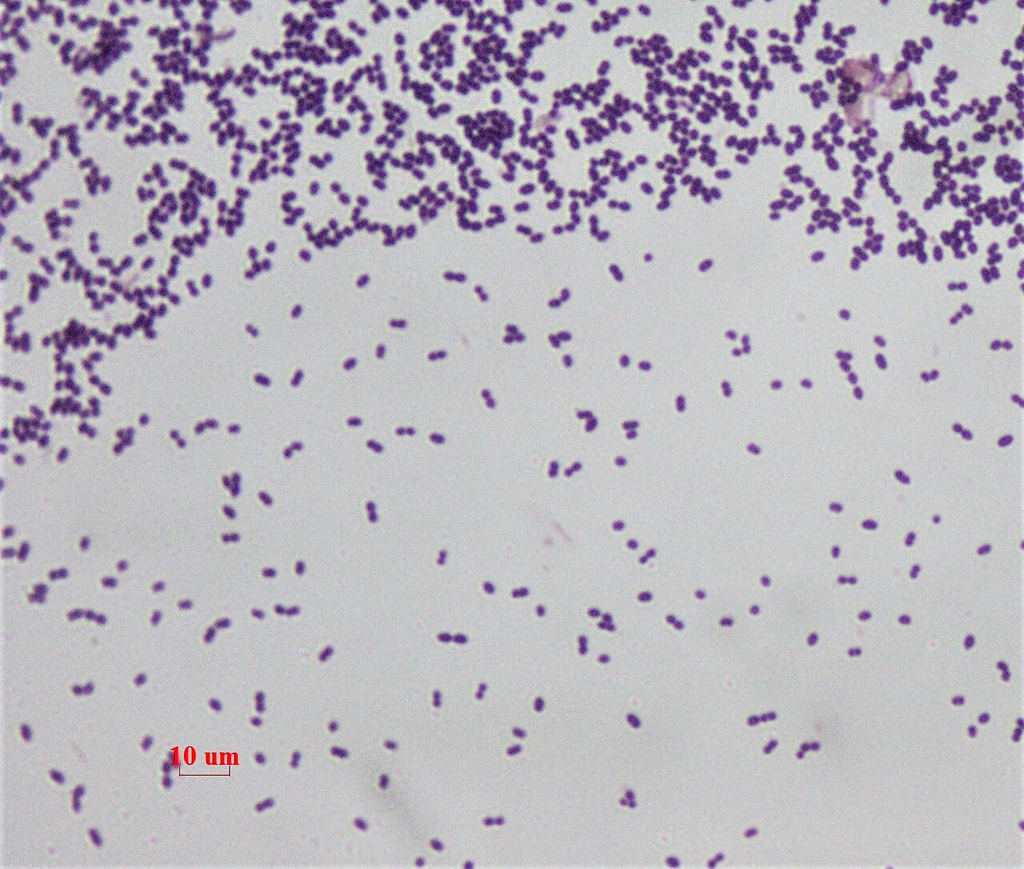
Gram stain of Enterococcus faecalis showing gram-positive cocci in pairs and short chains. Important nosocomial pathogen.
Oil immersion — diplococci and short chains
Wikimedia Commons — CC BY-SA 4.0
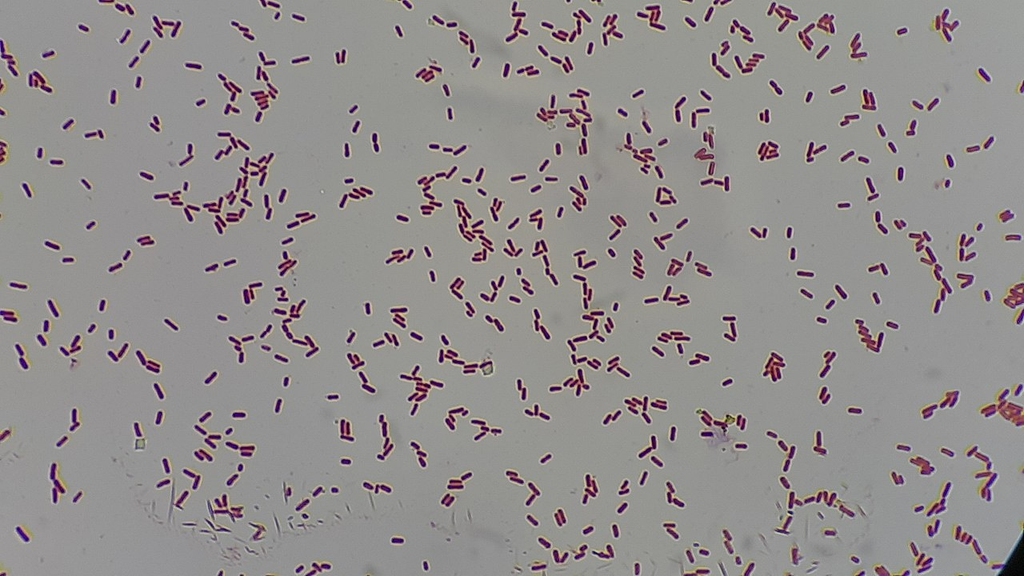
Gram stain of Klebsiella pneumoniae showing plump gram-negative bacilli. Mucoid capsule may appear as clear halo surrounding the organisms.
Oil immersion — encapsulated gram-negative rods
Wikimedia Commons — Public Domain (CDC)
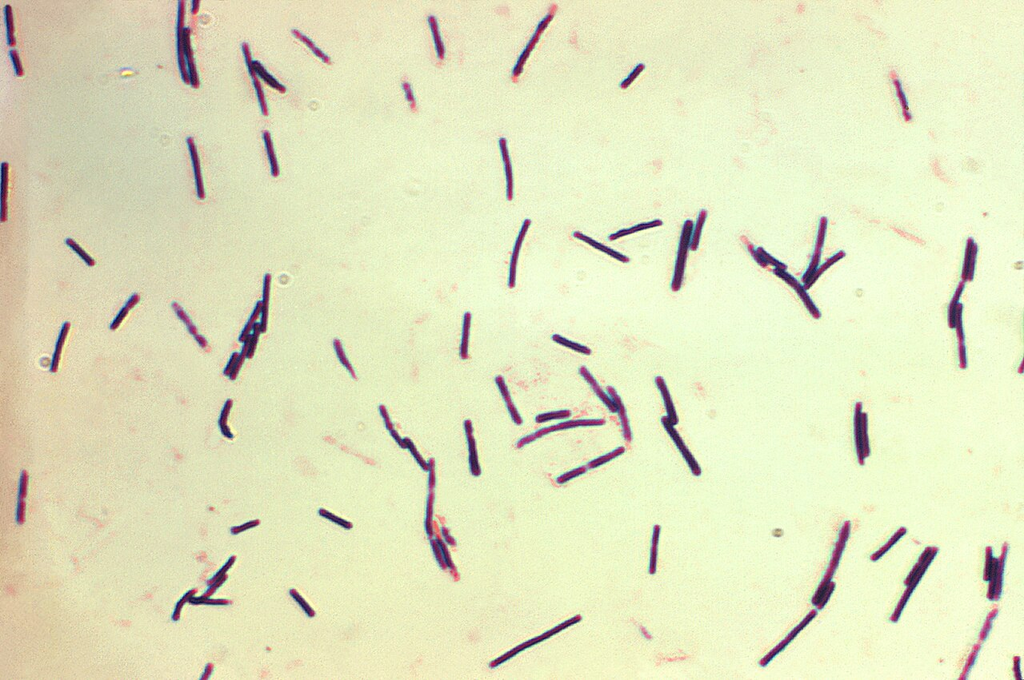
Gram stain of Clostridium perfringens showing large gram-positive bacilli. Characteristically large, box-car shaped rods causing gas gangrene.
Oil immersion — large box-car rods, anaerobic spore-former
Wikimedia Commons — Public Domain (CDC)
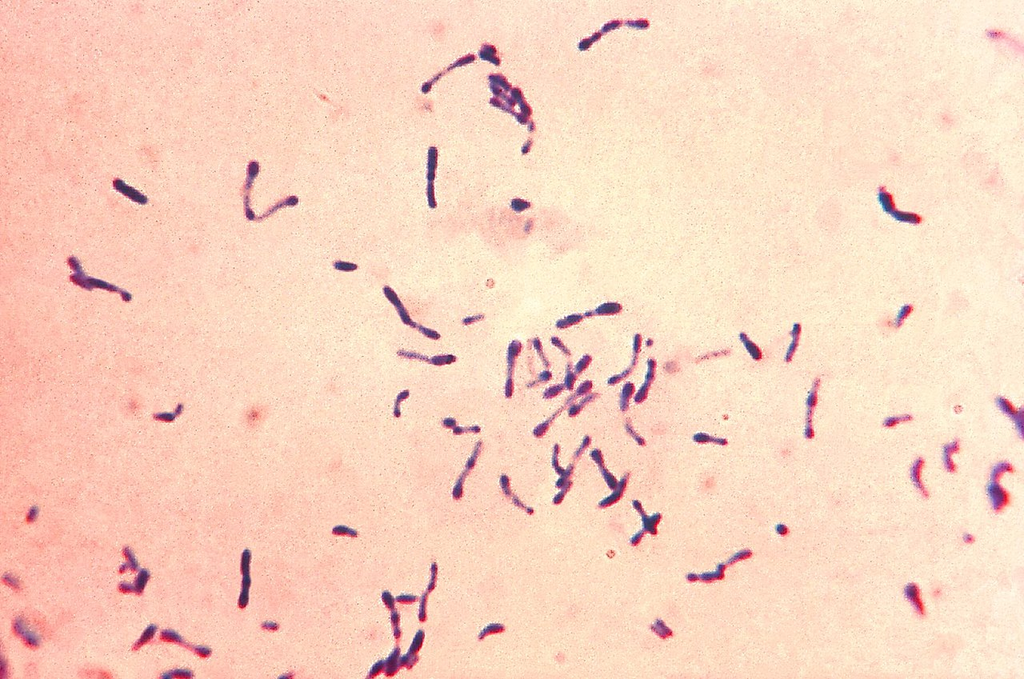
Gram-stained smear of Corynebacterium diphtheriae showing pleomorphic gram-positive bacilli in palisade and V/L arrangements (Chinese letter pattern).
Oil immersion — pleomorphic rods in palisade/Chinese letter arrangements
Wikimedia Commons — Public Domain (CDC)
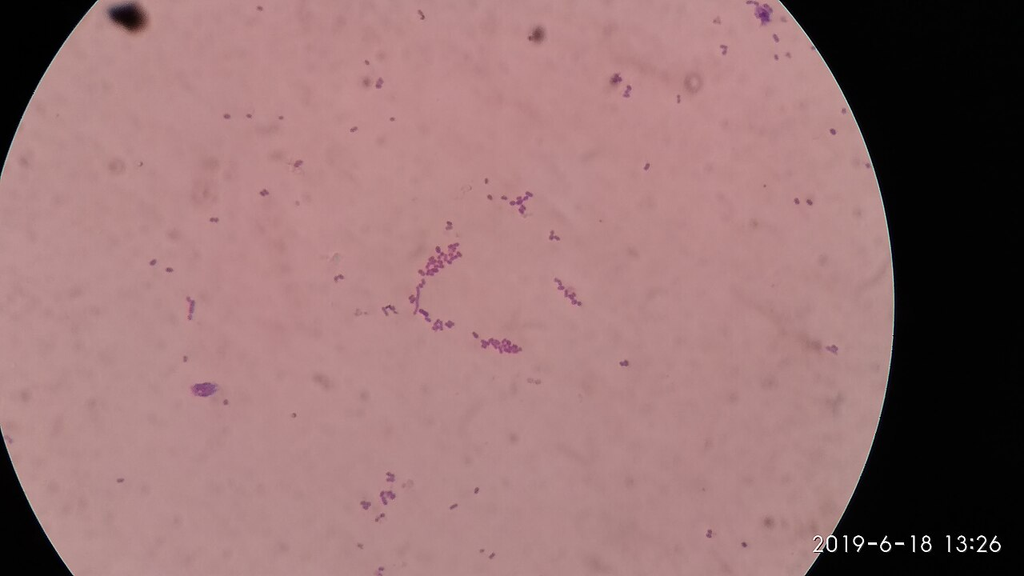
Gram stain of Listeria monocytogenes showing small gram-positive coccobacilli. Short rods that can resemble cocci, sometimes in short chains.
Oil immersion — small gram-positive coccobacilli, intracellular pathogen
Wikimedia Commons — CC BY-SA 3.0
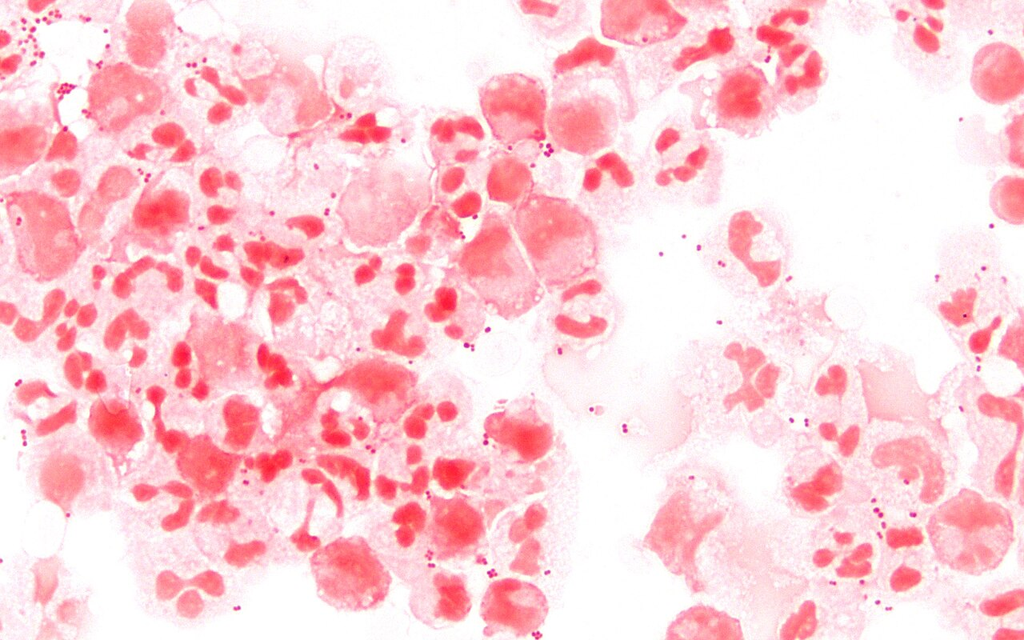
Gram stain of cerebrospinal fluid showing intracellular gram-negative diplococci within neutrophils, consistent with Neisseria meningitidis.
Oil immersion — intracellular gram-negative diplococci in CSF
Wikimedia Commons — Public Domain (CDC)
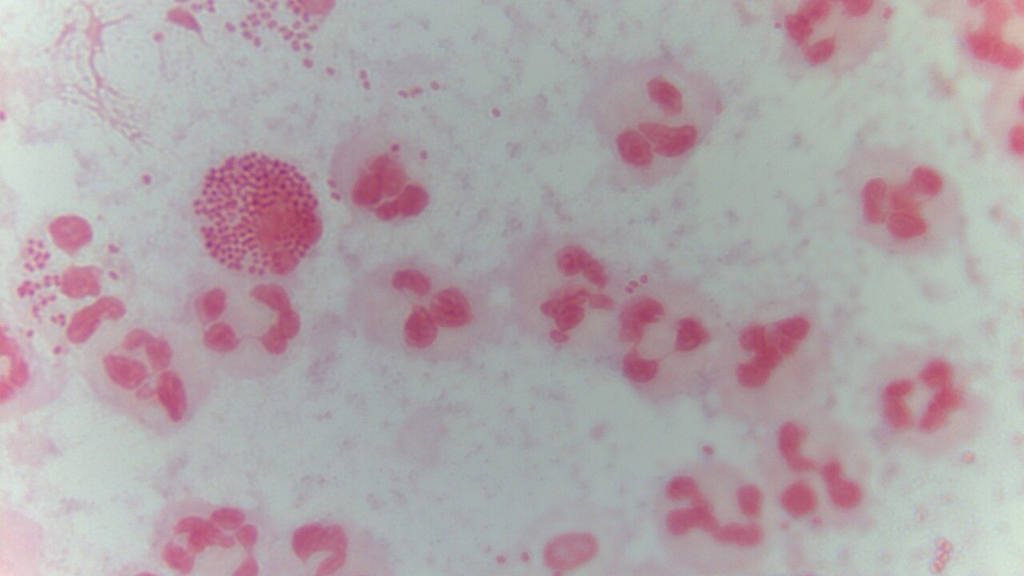
Gram-stained urethral discharge showing intracellular gram-negative diplococci within polymorphonuclear leukocytes. Classic finding for gonococcal urethritis.
Oil immersion — intracellular kidney-bean diplococci
Wikimedia Commons — Public Domain (CDC)
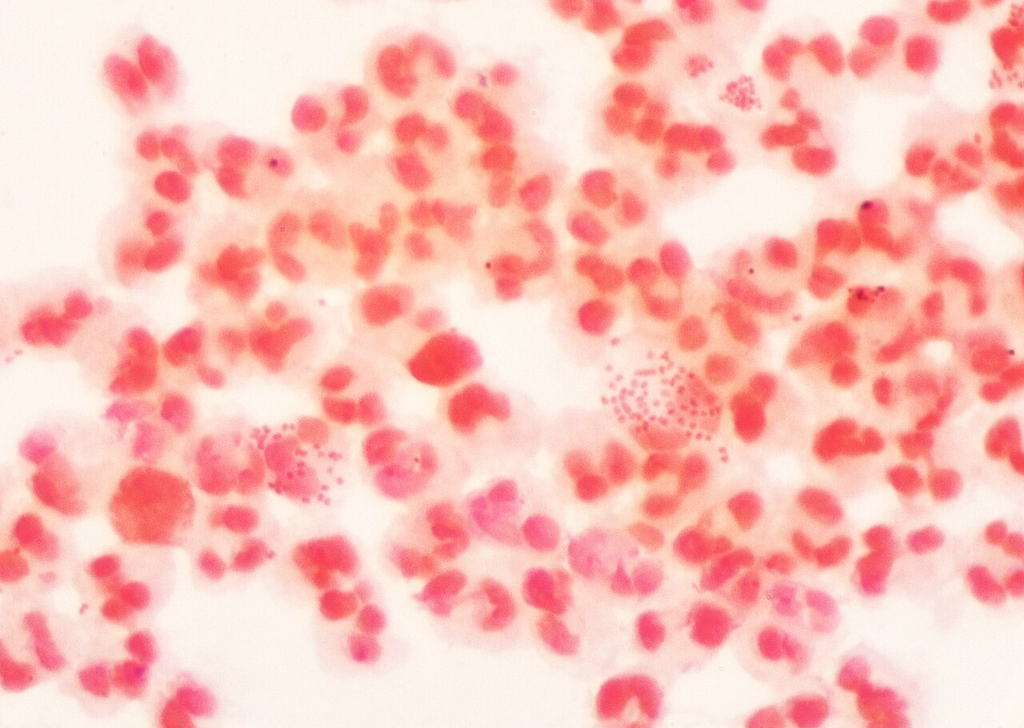
Gram stain showing gram-negative diplococci with numerous pus cells (PMNs). Characteristic finding in clinical specimen from gonococcal infection.
Oil immersion — gram-negative diplococci, respiratory pathogen
Wikimedia Commons — CC BY-SA 4.0
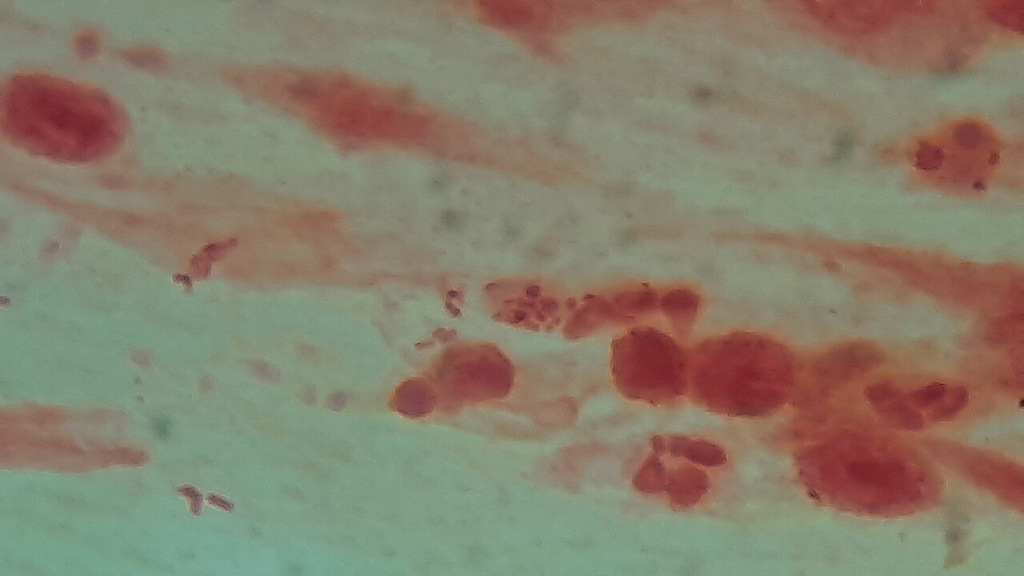
Gram-stained sputum specimen showing numerous gram-negative bacteria with polymorphonuclear leukocytes indicating lower respiratory infection.
Oil immersion — mixed flora with PMNs, lower respiratory specimen
Wikimedia Commons — CC BY-SA 3.0
Gram-stained smear of sputum prepared for microscopic observation showing mixed bacterial morphotypes and inflammatory cells.
Oil immersion — positive blood culture, cocci in clusters
Wikimedia Commons — CC BY-SA 3.0
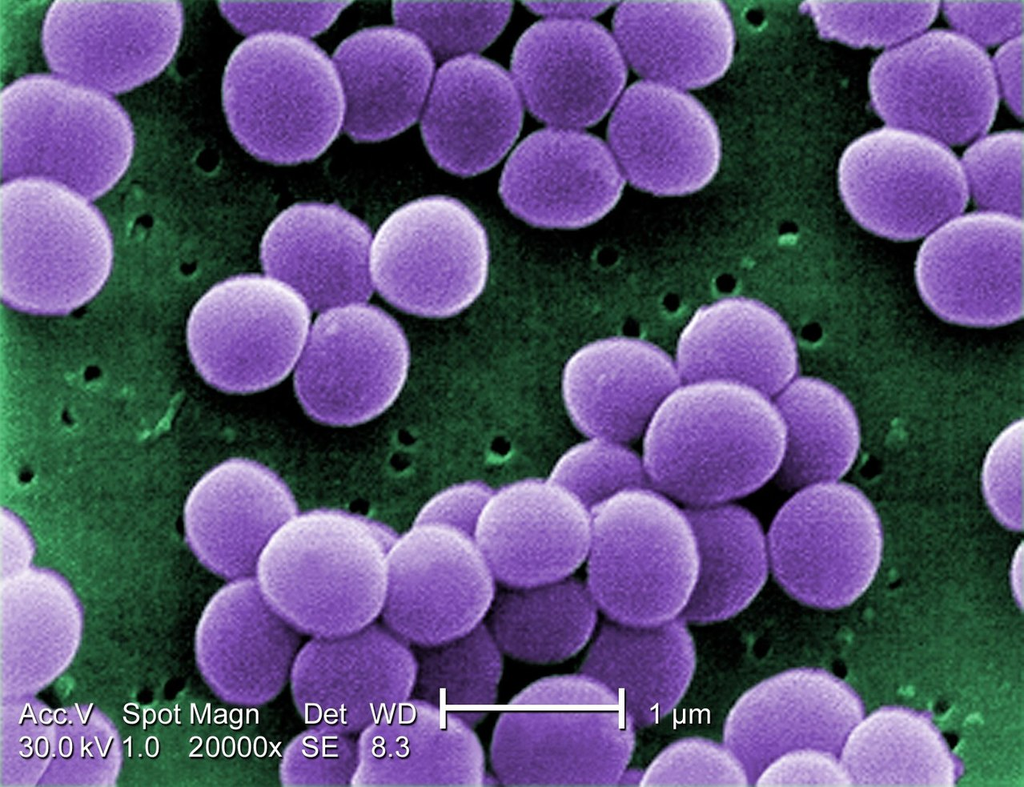
Transmission electron micrograph of Staphylococcus aureus (VISA strain) showing cell wall thickening, cell membrane, and internal cellular structures. Cross-section reveals thick peptidoglycan layer characteristic of gram-positive bacteria.
TEM thin section — cell wall thickening visible in VISA strain
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
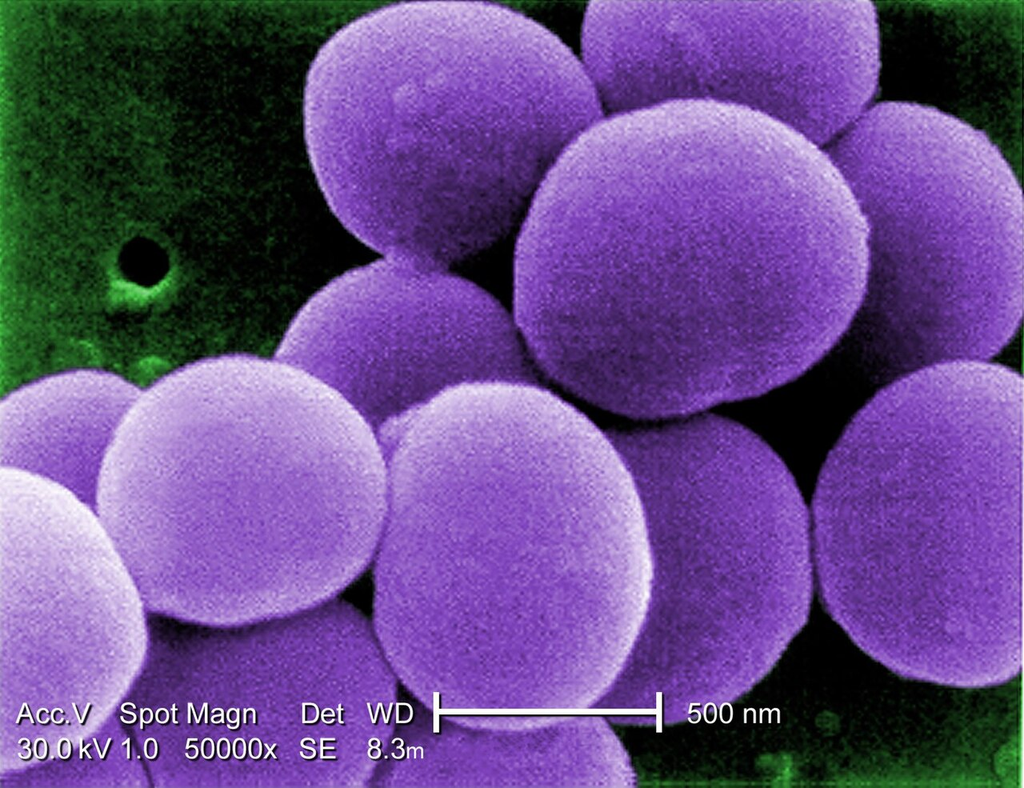
TEM cross-section of S. aureus (VISA strain) showing dividing cells with visible septum formation, thick peptidoglycan cell wall, cytoplasmic membrane, ribosomes, and nucleoid region.
TEM thin section — dividing cells with septum visible
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
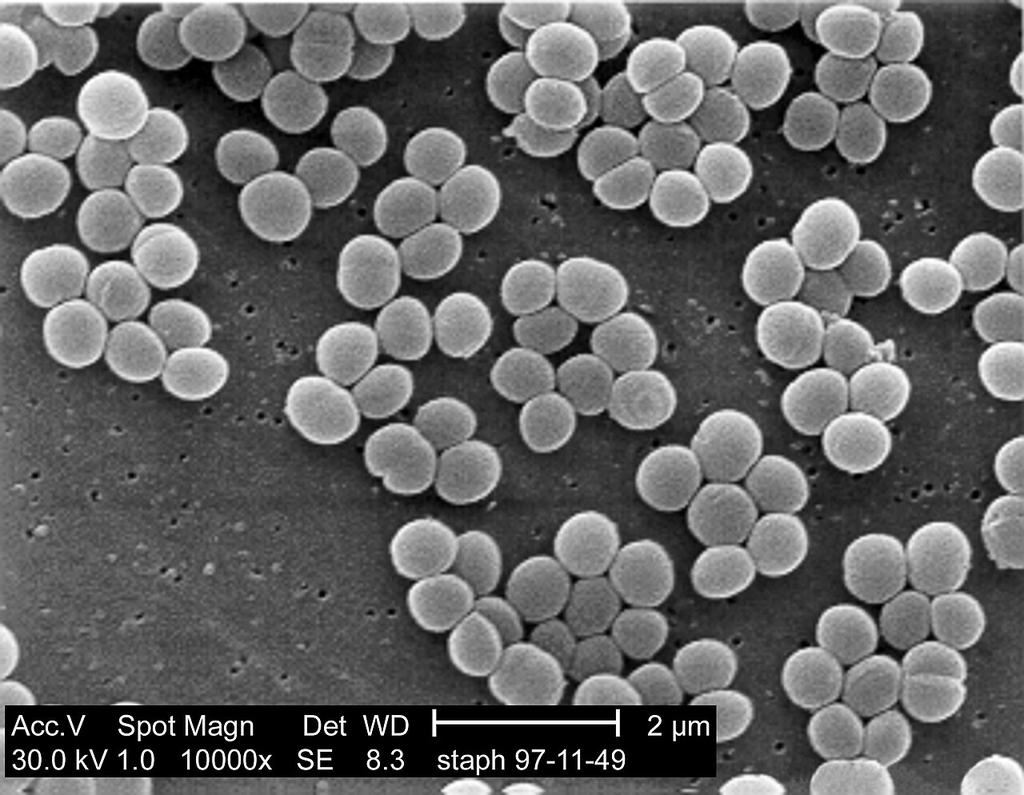
Electron micrograph of Staphylococcus aureus showing characteristic cocci morphology and surface detail at high magnification.
TEM — S. aureus surface and morphology detail
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
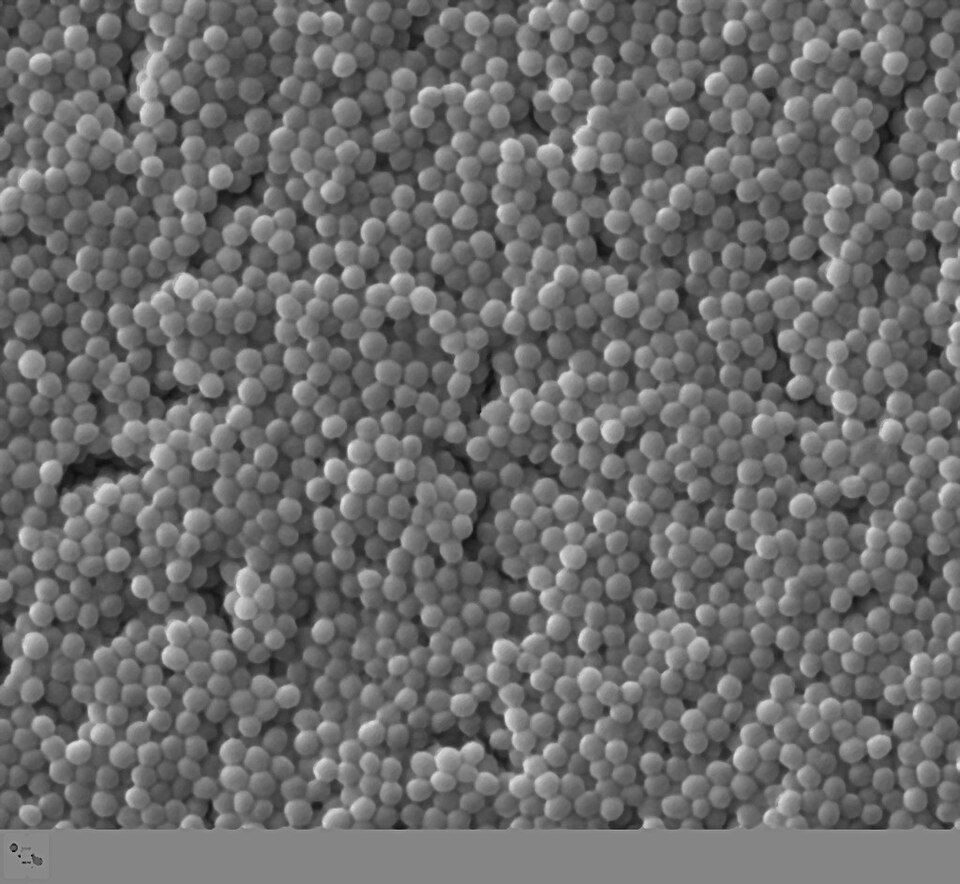

TEM image showing intact population of Staphylococcus aureus cells. Individual bacteria show cell wall, membrane, and cytoplasmic contents including ribosomes and nucleoid.
TEM — intact population showing internal structures
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
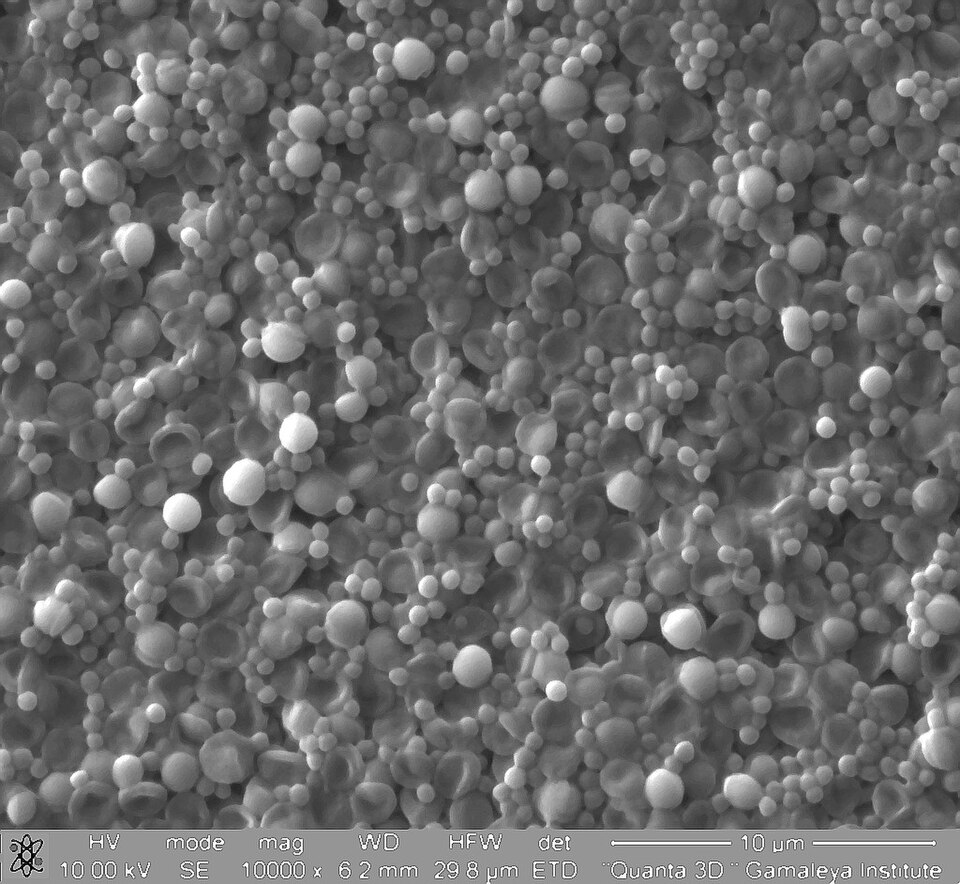

TEM image demonstrating phenotypic variation in S. aureus population. Cells show varying cell wall thickness and internal organization.
TEM — phenotypic variation in cell wall thickness
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
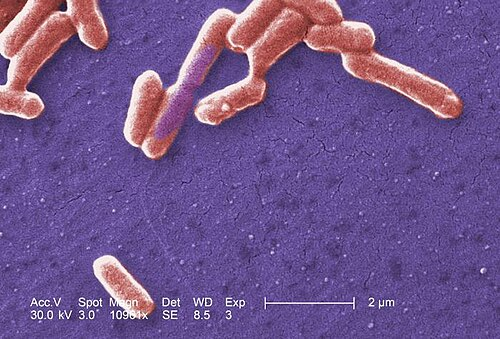
Scanning electron micrograph of an E. coli colony showing rod-shaped gram-negative bacteria. Surface detail reveals smooth outer membrane characteristic of Enterobacteriaceae.
SEM — E. coli colony surface morphology
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
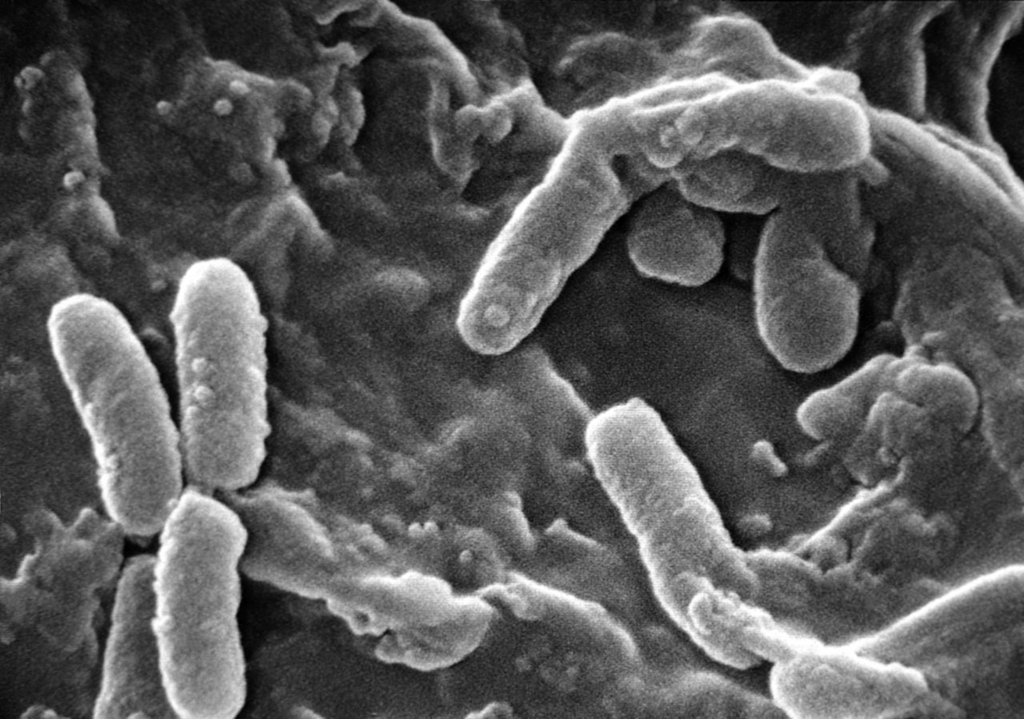
Scanning electron micrograph of Pseudomonas aeruginosa showing rod-shaped gram-negative bacteria with visible polar flagellum and surface pili.
SEM — P. aeruginosa with flagellum and pili visible
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
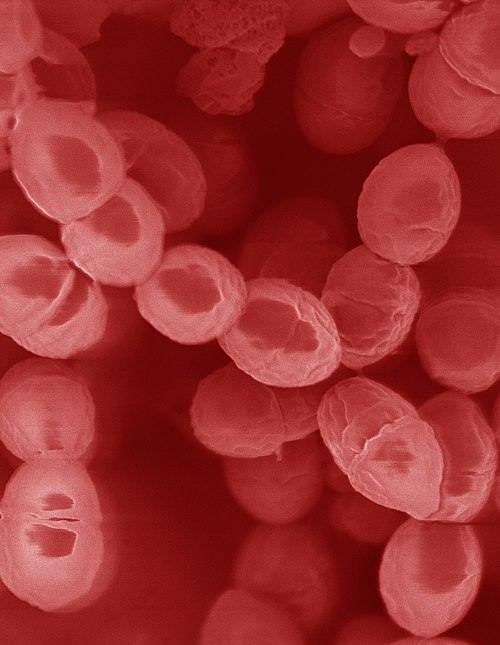
Scanning electron micrograph of Lactococcus lactis showing ovoid gram-positive cocci. Surface detail and cell division visible.
SEM — L. lactis cocci morphology and division
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
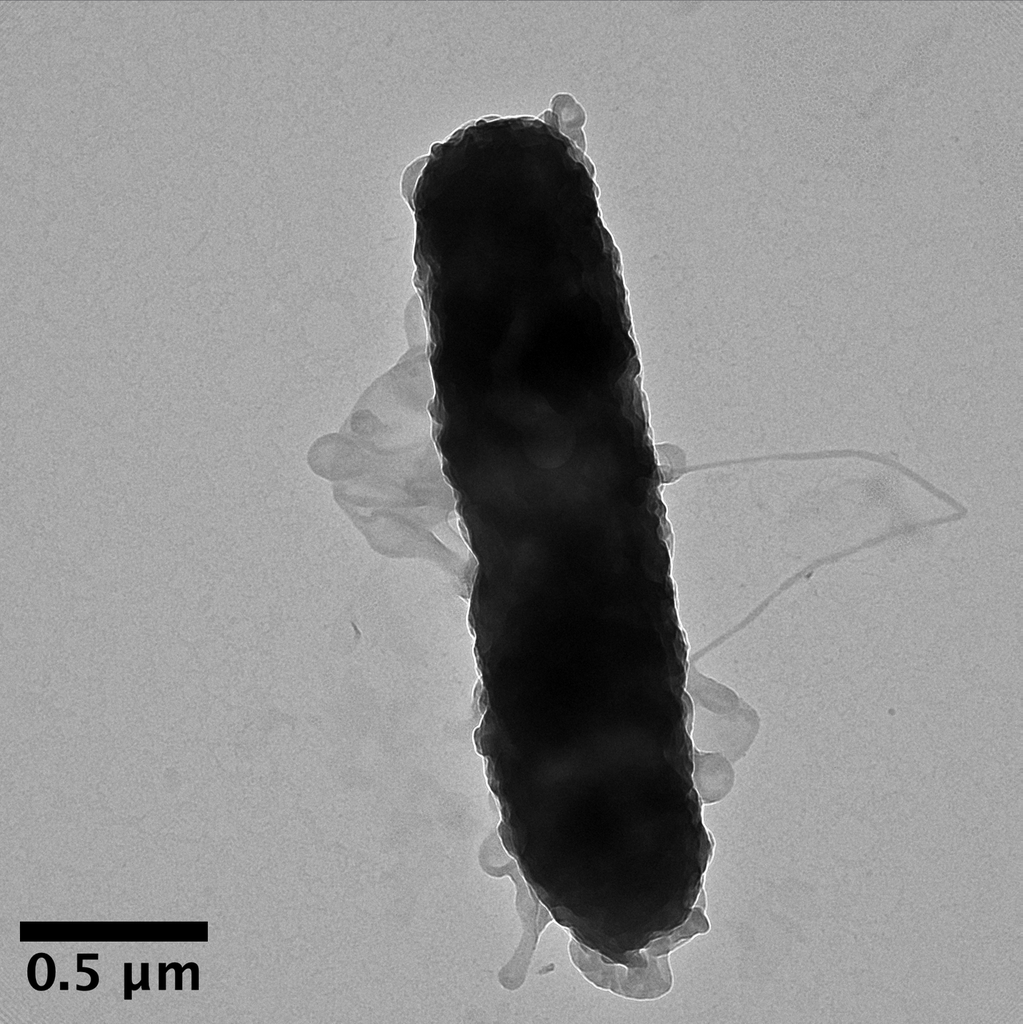

TEM image showing emerging projections of the cellular envelope of Pseudomonas sp. Outer membrane, periplasmic space, and inner membrane visible — classic gram-negative cell wall architecture.
TEM — gram-negative envelope architecture, outer membrane projections
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
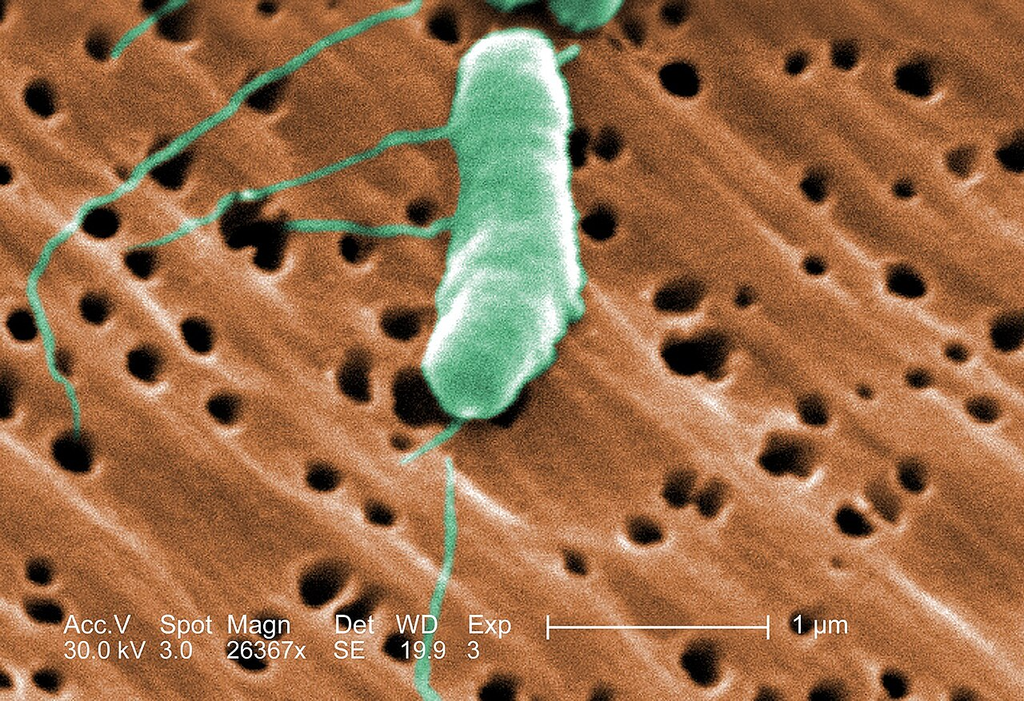
Colorized scanning electron micrograph of Vibrio vulnificus showing curved rod morphology with polar flagellum. Characteristic comma-shaped gram-negative bacterium.
SEM colorized — V. vulnificus curved rod with flagellum
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
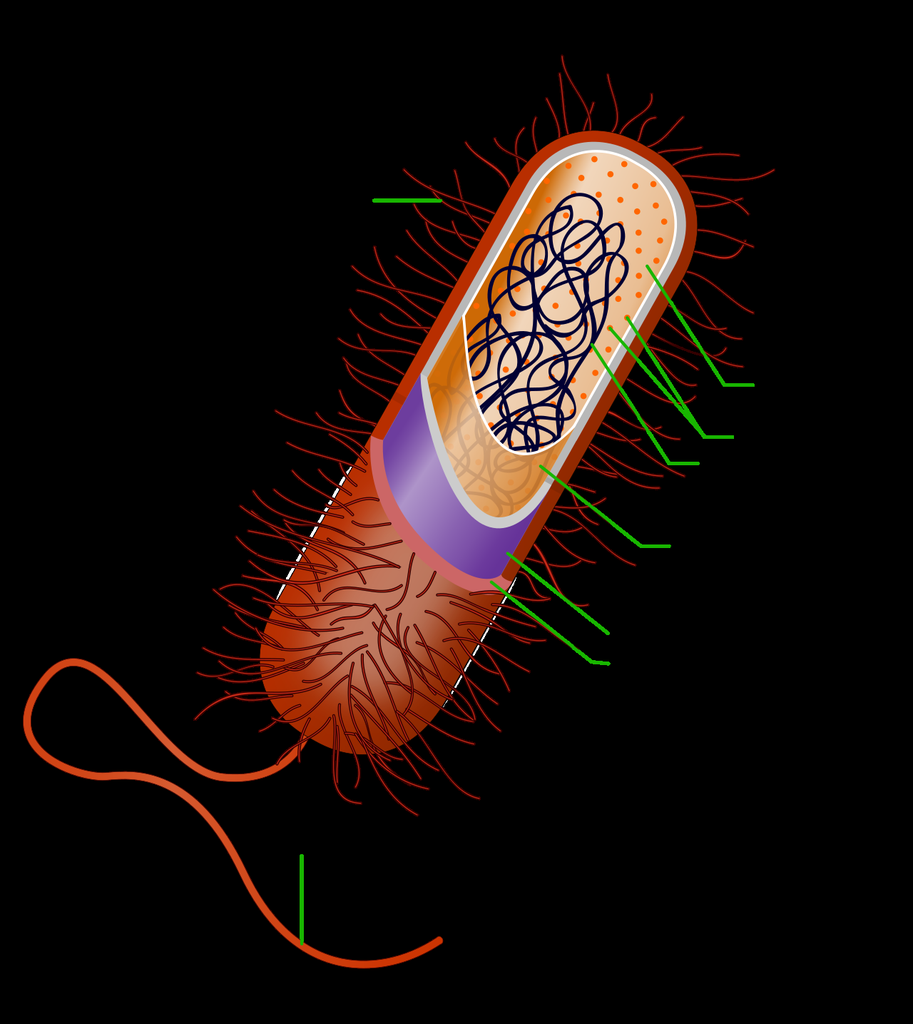
Labeled diagram of a prokaryotic cell showing cell wall, cell membrane, cytoplasm, ribosomes, nucleoid (DNA), flagellum, pili, and capsule.
Reference diagram — prokaryotic cell anatomy
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
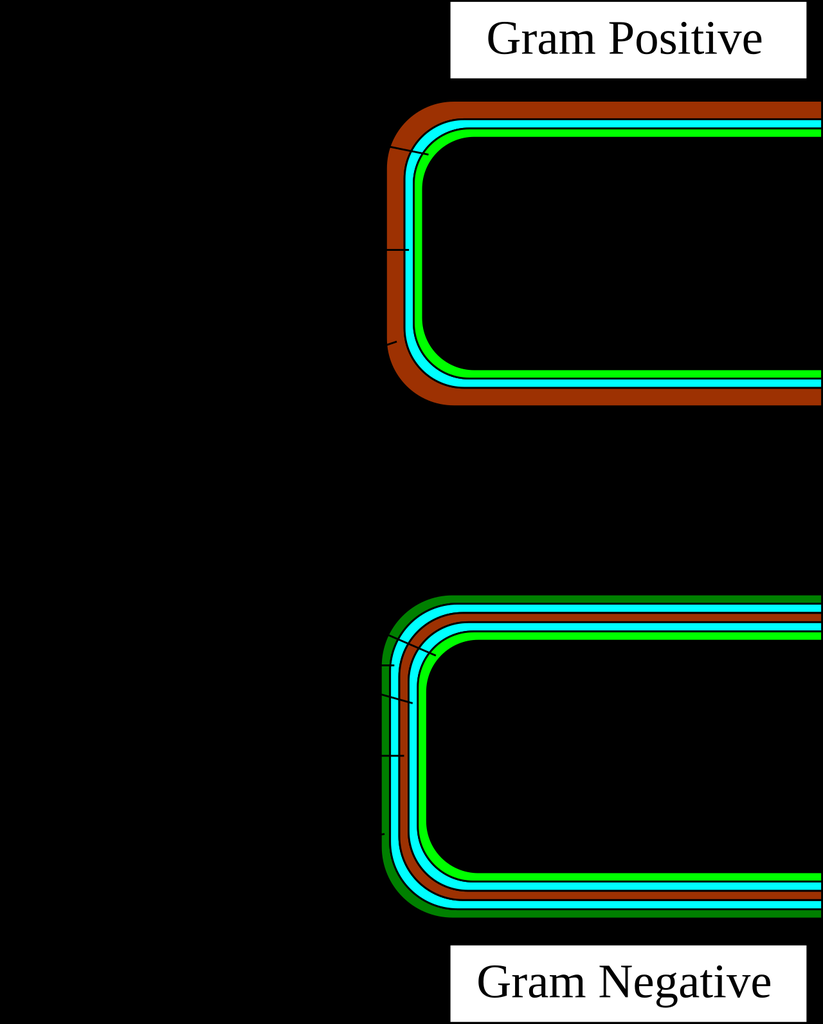

Comparative diagram showing gram-positive (thick peptidoglycan, teichoic acids) vs gram-negative (thin peptidoglycan, outer membrane with LPS, periplasmic space) cell wall architecture.
Reference diagram — comparative cell wall architecture
Wikimedia Commons — CC BY-SA 3.0 / Public Domain
Detailed labeled SVG diagram of a gram-positive bacterium (S. aureus) showing capsule, thick peptidoglycan cell wall, teichoic acids, cytoplasmic membrane, nucleoid, ribosomes (70S), plasmids, mesosome, pili/fimbriae, and cytoplasm. TEM cross-section perspective.
SVG diagram — labeled cross-section of gram-positive bacterial cell
Generated — Educational SVG
Detailed labeled SVG diagram of a gram-negative bacterium (E. coli) showing outer membrane, LPS (lipopolysaccharide), periplasmic space, thin peptidoglycan layer, inner membrane, nucleoid, ribosomes (70S), plasmids, flagellum, Type I pili, inclusion bodies, and cytoplasm. TEM cross-section perspective.
SVG diagram — labeled cross-section of gram-negative bacterial cell
Generated — Educational SVG
Side-by-side comparative SVG diagram showing gram-positive (thick peptidoglycan, teichoic acids) vs gram-negative (thin peptidoglycan, outer membrane, LPS, periplasmic space, porins) cell wall architecture. Includes dimensional annotations.
SVG diagram — comparative cell wall architecture at high TEM magnification
Generated — Educational SVG
Compatible Microscopes
| Model | Manufacturer | Type | Magnification Range | NA Max | Resolution |
|---|---|---|---|---|---|
| HT7800 | Hitachi High-Tech | Tem | 200–600000× | — | — nm |
| Regulus 8200 | Hitachi High-Tech | Fe Sem | 20–2000000× | — | 0.6 nm |
| JEM-F200 | JEOL | Tem | 50–1500000× | — | — nm |
| JSM-IT800 | JEOL | Sem | 5–1000000× | — | 0.7 nm |
| JSM-7900F | JEOL | Fe Sem | 25–1000000× | — | 0.5 nm |
| DM6 B | Leica Microsystems | Upright Optical | 1.25–100× | 1.4 | 200 nm |
| DMi8 | Leica Microsystems | Inverted Optical | 2.5–100× | 1.47 | 185 nm |
| Eclipse Ti2 | Nikon | Inverted Optical | 2–100× | 1.45 | 190 nm |
| Eclipse Ni | Nikon | Upright Optical | 2–100× | 1.4 | 200 nm |
| BX53 | Olympus (Evident) | Upright Optical | 2–100× | 1.4 | 200 nm |
| IX83 | Olympus (Evident) | Inverted Optical | 2–100× | 1.4 | 200 nm |
| Apreo 2 | Thermo Fisher Scientific | Fe Sem | 13–2000000× | — | 0.7 nm |
| Axio Observer 7 | Zeiss | Inverted Optical | 5–100× | 1.4 | 200 nm |
| Axio Imager 2 | Zeiss | Upright Optical | 1.25–100× | 1.4 | 200 nm |
| Primostar 3 | Zeiss | Upright Optical | 4–100× | 1.25 | 350 nm |