AI Brain Tissue (Neurons)
AI Brain Tissue (Neurons)
NeuroscienceAI-generated brain tissue image set spanning the full range of microscopy magnification — from 4× scanning power through 100,000× transmission electron microscopy. Each image is generated at a specific magnification increment with the appropriate microscope, stain, and optical parameters for that level. Includes Nissl-stained, Golgi-stained, H&E-stained, and unstained ultrastructural views of cerebral cortex neurons, glial cells, and neuropil. Designed as a continuous zoom-through training reference for neuroanatomy and neuropathology education.
Drag & drop images here to add to this specimen
or click Upload Image aboveRelease to upload
Image Library
12 images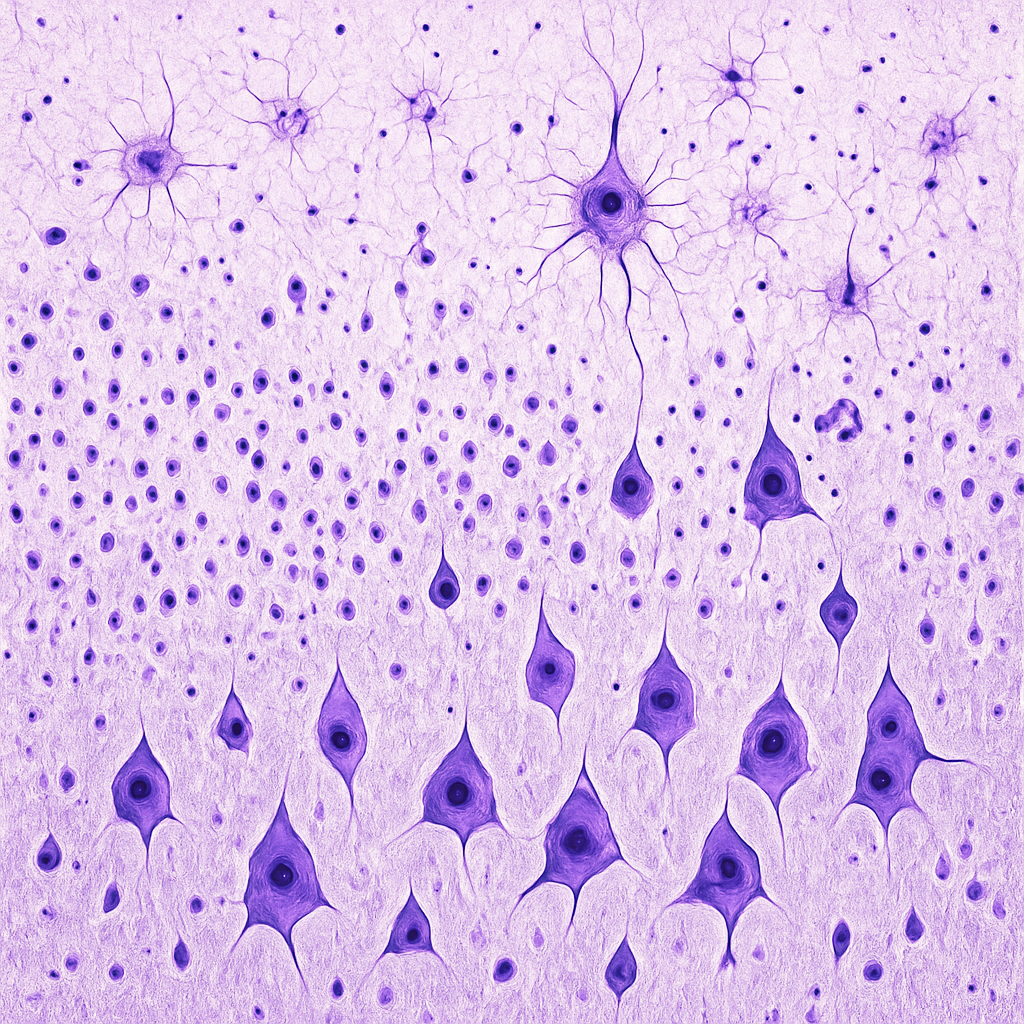
Brain tissue section at 4× scanning objective showing the full thickness of cerebral cortex. Nissl-stained (cresyl violet) section reveals the six cortical layers (I–VI) distinguished by neuron size, density, and packing pattern. Layer I (molecular layer) appears pale with sparse cells. Layers II/III (external pyramidal) show densely packed small-to-medium neurons. Layer IV (internal granular) has many small stellate cells. Layer V (internal pyramidal) contains large pyramidal neurons (Betz cells). Layer VI (polymorphic) transitions to the white matter below.
4× scanning objective, NA 0.1. Nissl stain (cresyl violet) highlights neuronal cell bodies and rough endoplasmic reticulum. Used to identify cortical layers and cytoarchitectural regions. Zeiss Primostar 3 — teaching-grade clinical microscope.
OpenAI gpt-image-1
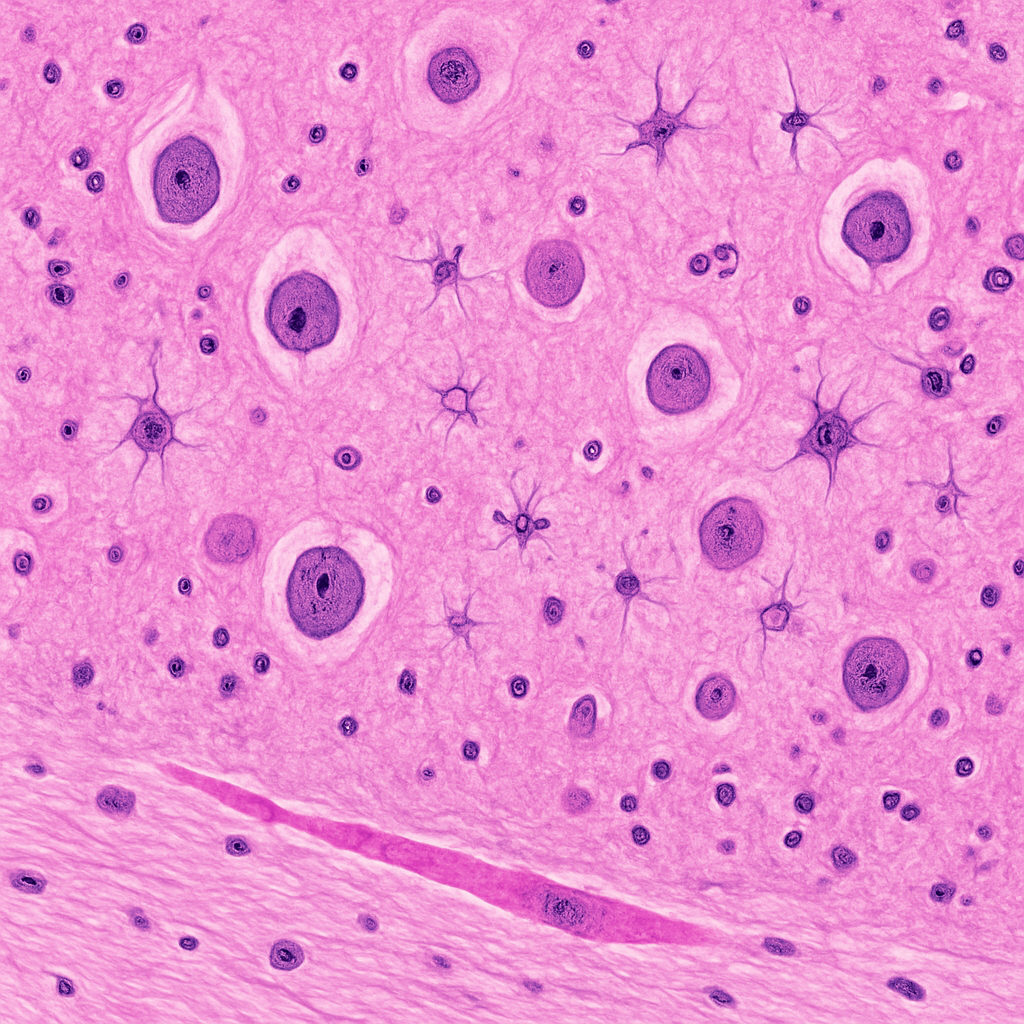
H&E-stained cerebral cortex section at 10× low-power objective. The neuropil appears as a pink fibrillar meshwork. Neuronal cell bodies are scattered throughout with pale round nuclei and prominent nucleoli. The gray matter shows a mix of neurons and smaller glial cells. Blood vessels with pink-stained walls are visible cutting through the cortical parenchyma. The boundary between gray and white matter is visible at the bottom of the field, with the white matter showing parallel myelinated fiber bundles.
10× objective, NA 0.25. H&E staining — nuclei blue-purple, cytoplasm and neuropil pink. Cortical architecture and gray-white matter boundary visible. Zeiss Axio Imager 2 — research-grade upright microscope.
OpenAI gpt-image-1
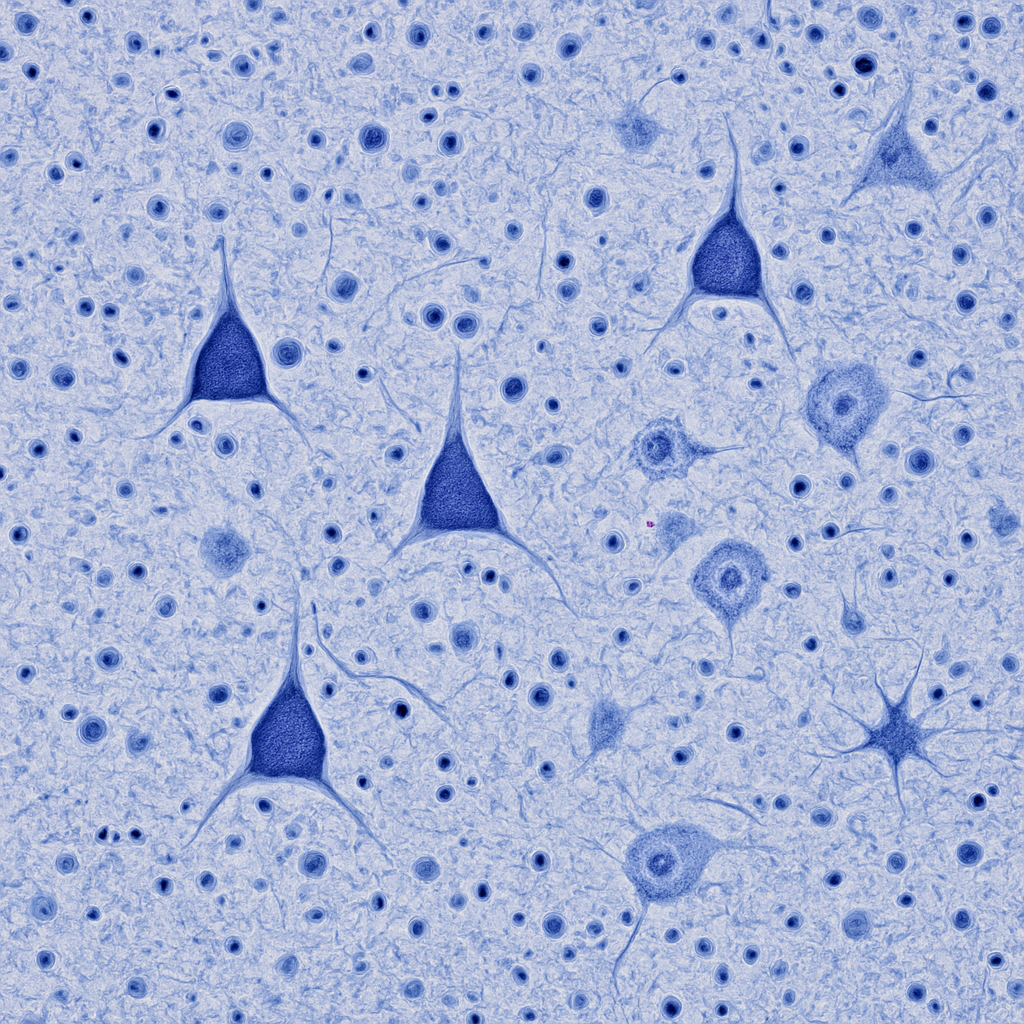

Nissl-stained cortex at 20× showing layers III and V with pyramidal neurons clearly visible. The pyramidal cells have triangular cell bodies stained dark blue-violet, with the apex pointing toward the cortical surface. Surrounding them is the pale neuropil interspersed with smaller glial cell nuclei (oligodendrocytes appearing as small dark dots, astrocytes as medium-sized pale nuclei). Neuropil — the tangled mesh of dendrites, axons, and synapses — fills the space between cell bodies as a pale blue-gray background.
20× objective, NA 0.40. Nissl stain reveals neuronal populations — pyramidal cells dominant in layers III/V. Glial nuclei distinguishable by size and shape. Olympus BX53 — clinical-grade upright microscope.
OpenAI gpt-image-1
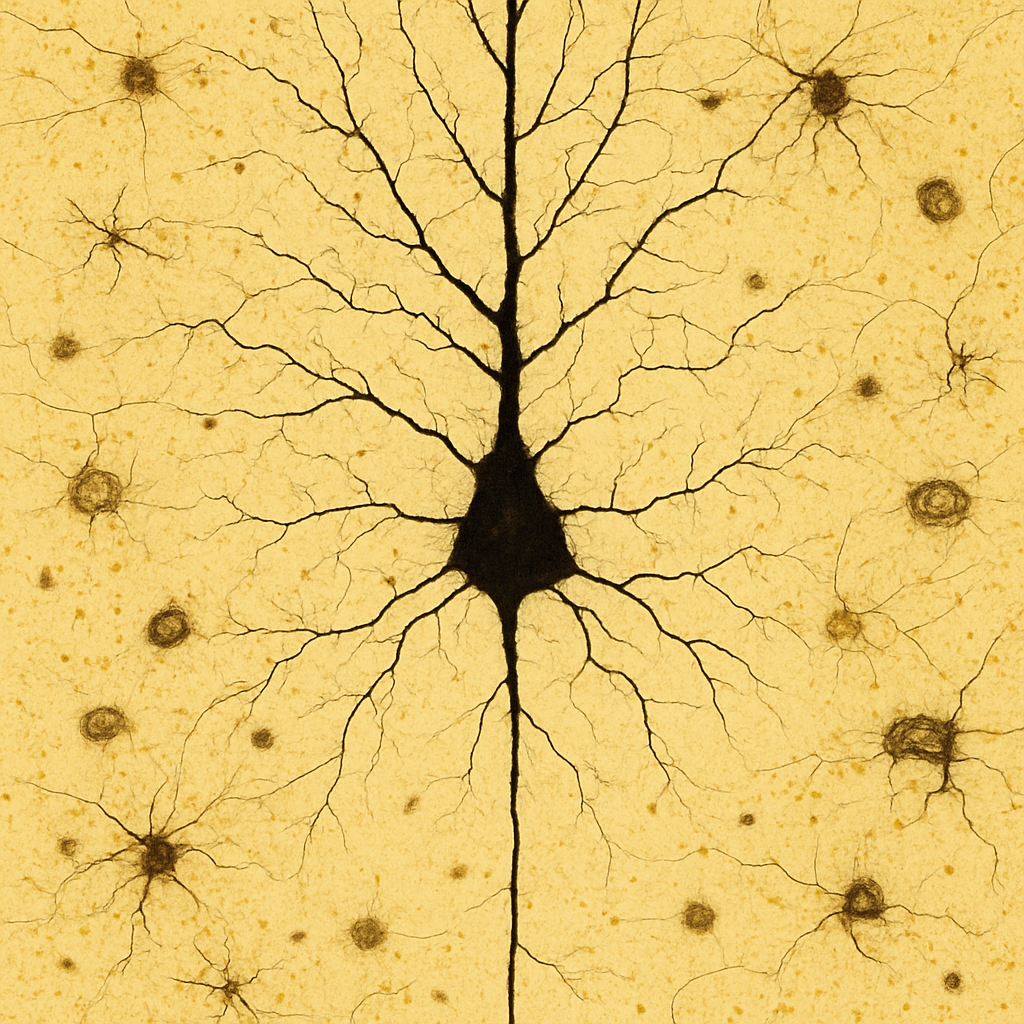
Golgi-stained cerebral cortex at 40× high-dry objective showing a single large pyramidal neuron with its complete dendritic arbor. The cell body (soma) is triangular, approximately 20 μm in diameter, with a prominent apical dendrite extending toward the cortical surface and multiple basal dendrites radiating from the base. The dendrites branch extensively and are studded with tiny dendritic spines — the sites of excitatory synaptic input. The thin axon extends downward from the base of the soma. Background neurons are partially stained. The Golgi method stains only 1–5% of neurons completely, rendering them in dark brown-black against a transparent golden-yellow background.
40× high-dry objective, NA 0.65. Golgi silver chromate stain — complete neuronal morphology visible including dendritic arbor and spines. Only ~1–5% of neurons are stained (random Golgi impregnation). Nikon Eclipse Ni — research-grade upright microscope.
OpenAI gpt-image-1
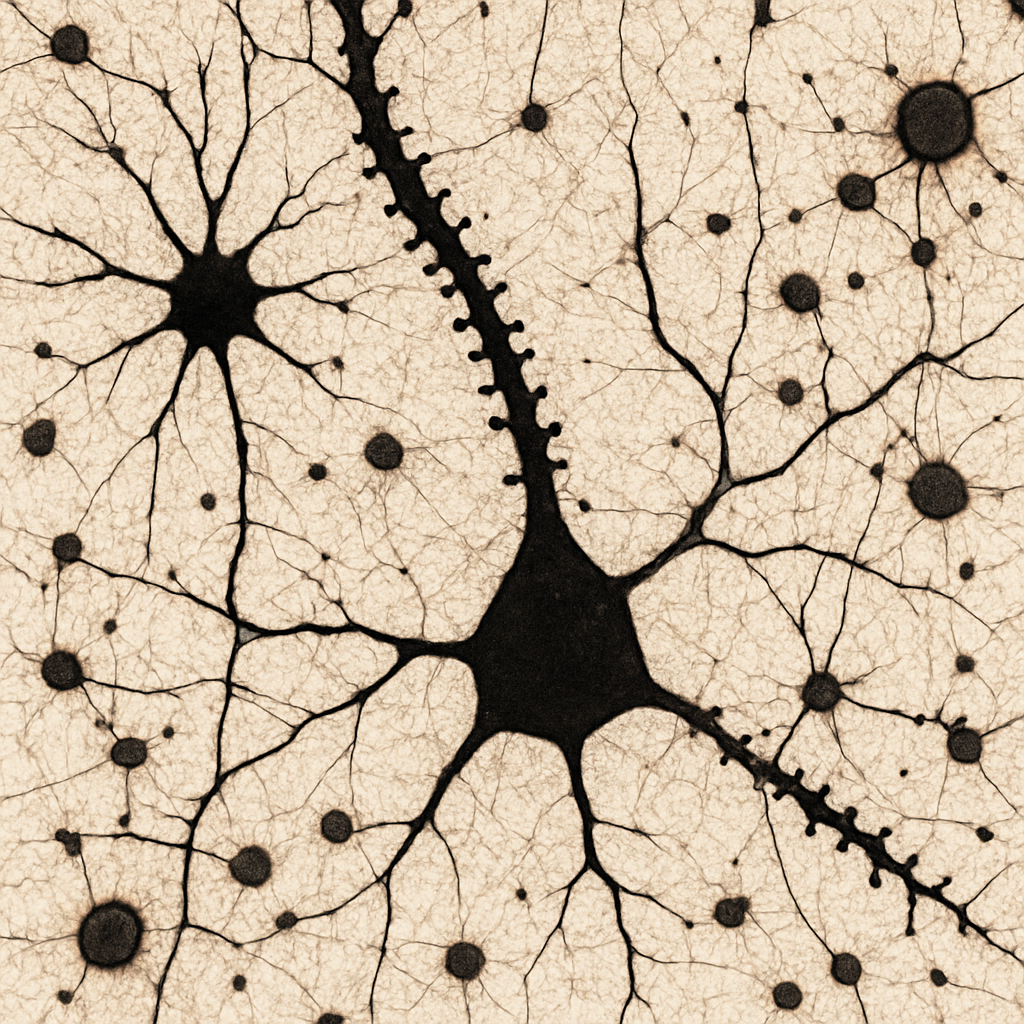
Golgi-stained section at 60× oil immersion showing a pyramidal neuron dendrite in fine detail. Individual dendritic spines are clearly resolved as small mushroom-shaped, thin, or stubby protrusions (1–2 μm) along the dendrite shaft. The spine heads — sites of glutamatergic synaptic input — are visible as tiny knobs. Near the dendrite, a Golgi-stained astrocyte is visible with its star-shaped morphology and fine processes extending toward nearby blood vessels (astrocytic endfeet). The fine fibrous processes of the neuropil create a delicate web-like background.
60× oil immersion, NA 1.25. Golgi stain — dendritic spines individually resolved. Astrocyte morphology with vascular endfeet visible. Leica DM6 B — research-grade upright with motorized stage.
OpenAI gpt-image-1
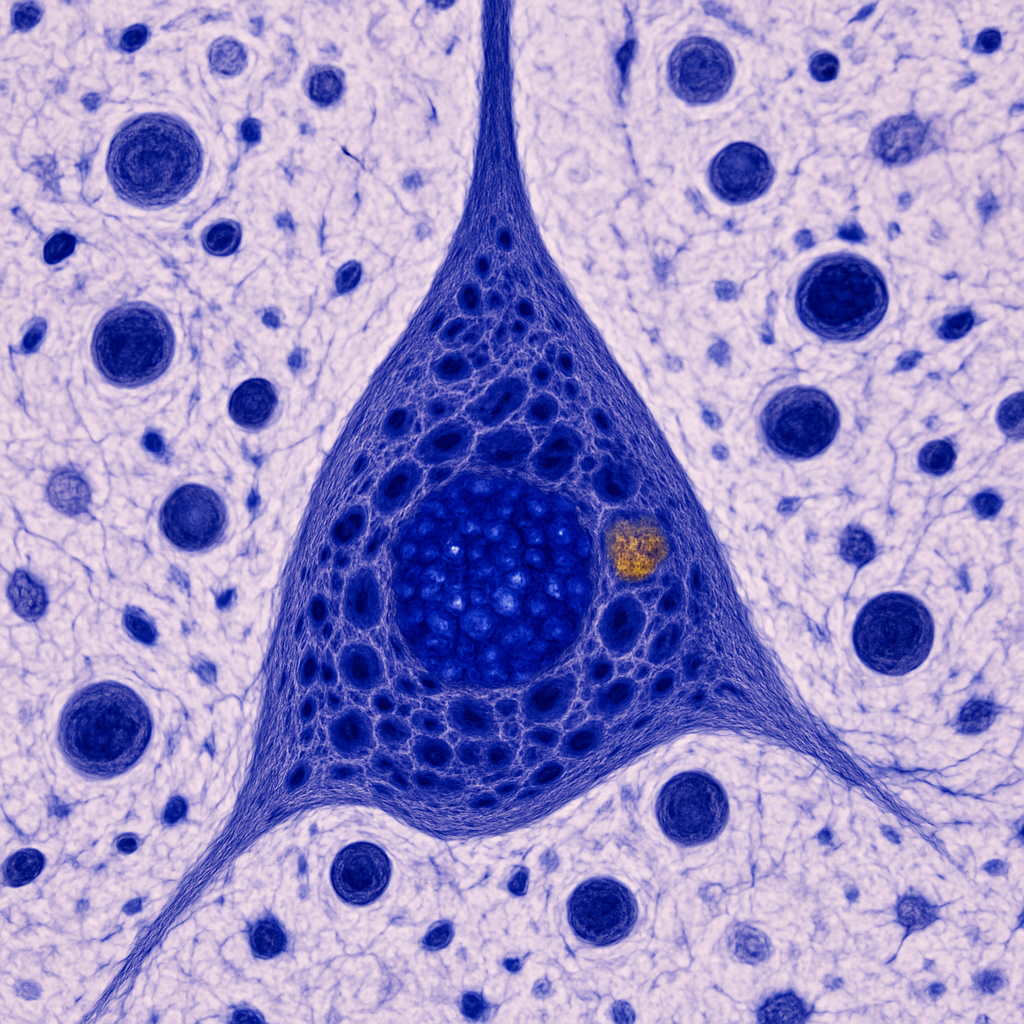
Nissl-stained brain at 100× oil immersion — maximum light microscopy resolution. A single large pyramidal neuron fills the field. The nucleus is large, round, and pale (euchromatic) with a single prominent dark nucleolus. The cytoplasm is filled with dark blue-purple Nissl substance (rough endoplasmic reticulum studded with ribosomes) arranged in large clumps called Nissl bodies or tigroid substance. The apical dendrite emerges from the apex of the triangular soma. A lipofuscin granule — a golden-brown age-related inclusion — may be visible in the cytoplasm. Surrounding the neuron, satellite oligodendrocytes appear as small dark round nuclei closely apposed to the neuronal cell body.
100× oil immersion, NA 1.40. Maximum resolving power for light microscopy (200 nm resolution). Nissl bodies, nucleolus, lipofuscin granules, and satellite oligodendrocytes all resolvable. Zeiss Axio Imager 2 — plan-apochromatic 100×/1.40 oil objective.
OpenAI gpt-image-1
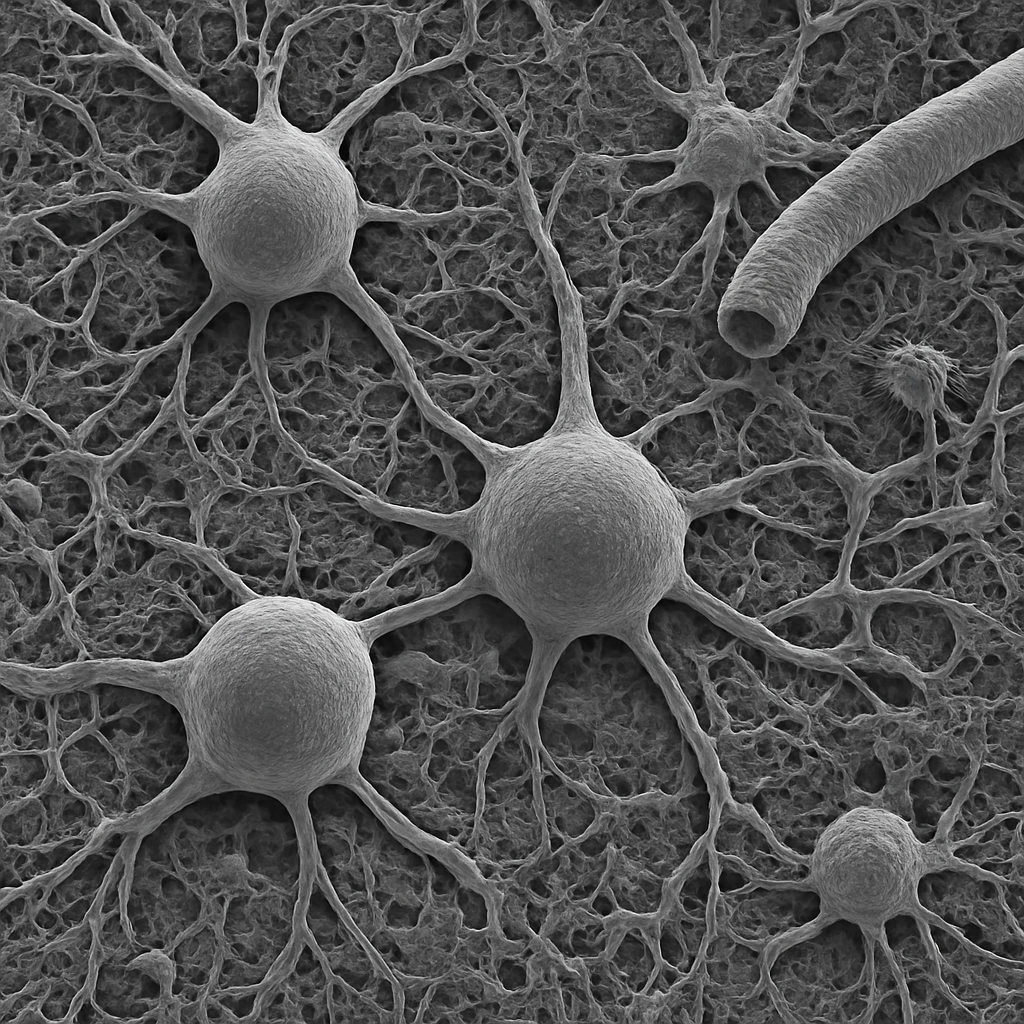
Scanning electron micrograph of brain tissue surface at 1,000× magnification. The cortical surface shows the complex three-dimensional topography of neuronal cell bodies, dendrites, and surrounding neuropil. Large pyramidal neuron somata appear as rounded elevations (~20 μm) with thick dendritic trunks extending from their apices. The neuropil between cell bodies is a dense fibrous mat of intertwined processes. Capillary blood vessels wind through the tissue as cylindrical channels. The image is grayscale with the characteristic 3D relief of secondary electron SEM.
1,000× SEM, secondary electron imaging. Osmium-fixed, critical-point dried, gold-palladium coated. Surface topography shows neuronal somata and neuropil architecture. JEOL JSM-IT800 — Schottky FEG, 0.7 nm res.
OpenAI gpt-image-1
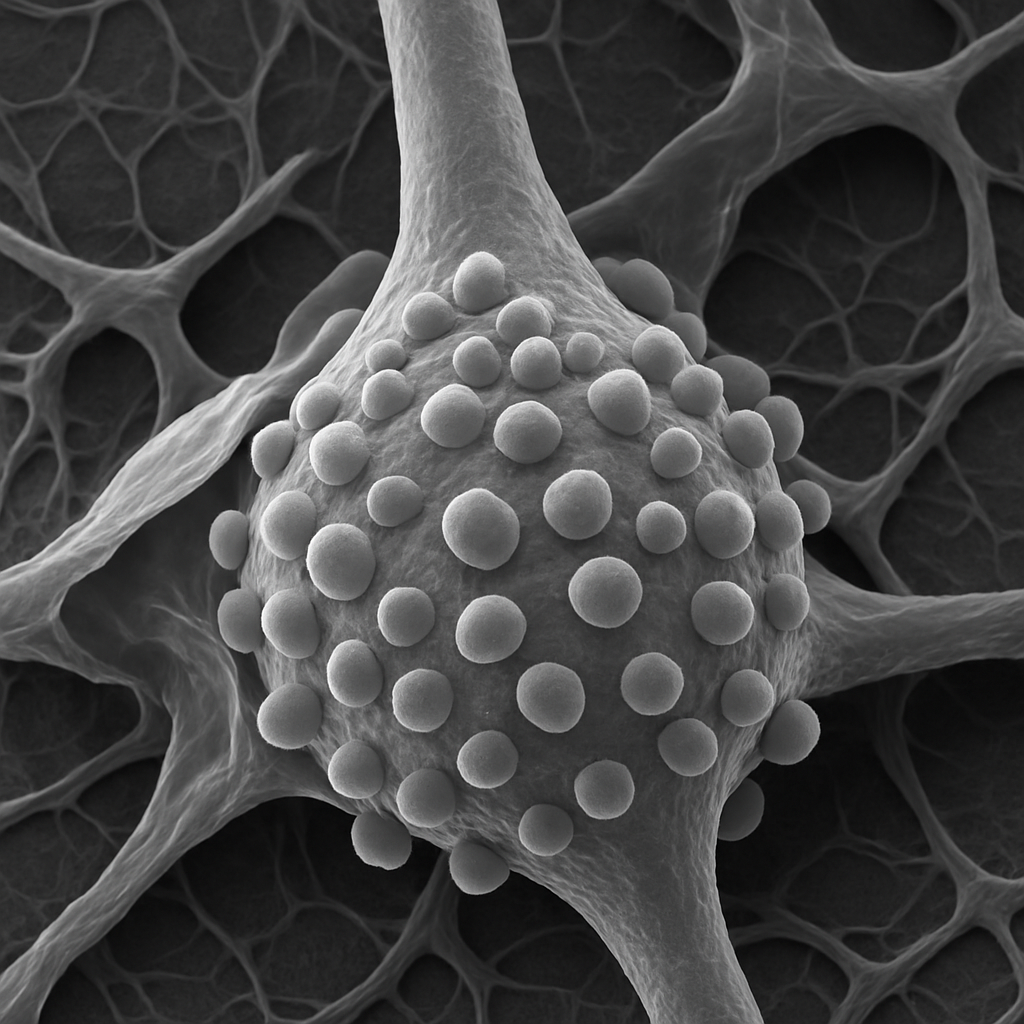
SEM at 5,000× showing the surface of a single pyramidal neuron soma and proximal dendrite. The cell body surface is covered with numerous synaptic boutons — small rounded presynaptic terminals (~1–2 μm) that contact the neuronal surface. These axosomatic synapses appear as hemisphere-like bumps pressed against the postsynaptic cell membrane. The proximal apical dendrite shows a smooth surface with occasional bouton contacts. Fine glial cell processes (astrocyte lamellae) wrap around the neuron surface in places, appearing as thin sheet-like coverings.
5,000× FE-SEM, secondary electron detector. Synaptic boutons, glial wrapping, and neuronal surface topology resolved. Hitachi Regulus 8200 — 0.6 nm resolution FE-SEM.
OpenAI gpt-image-1
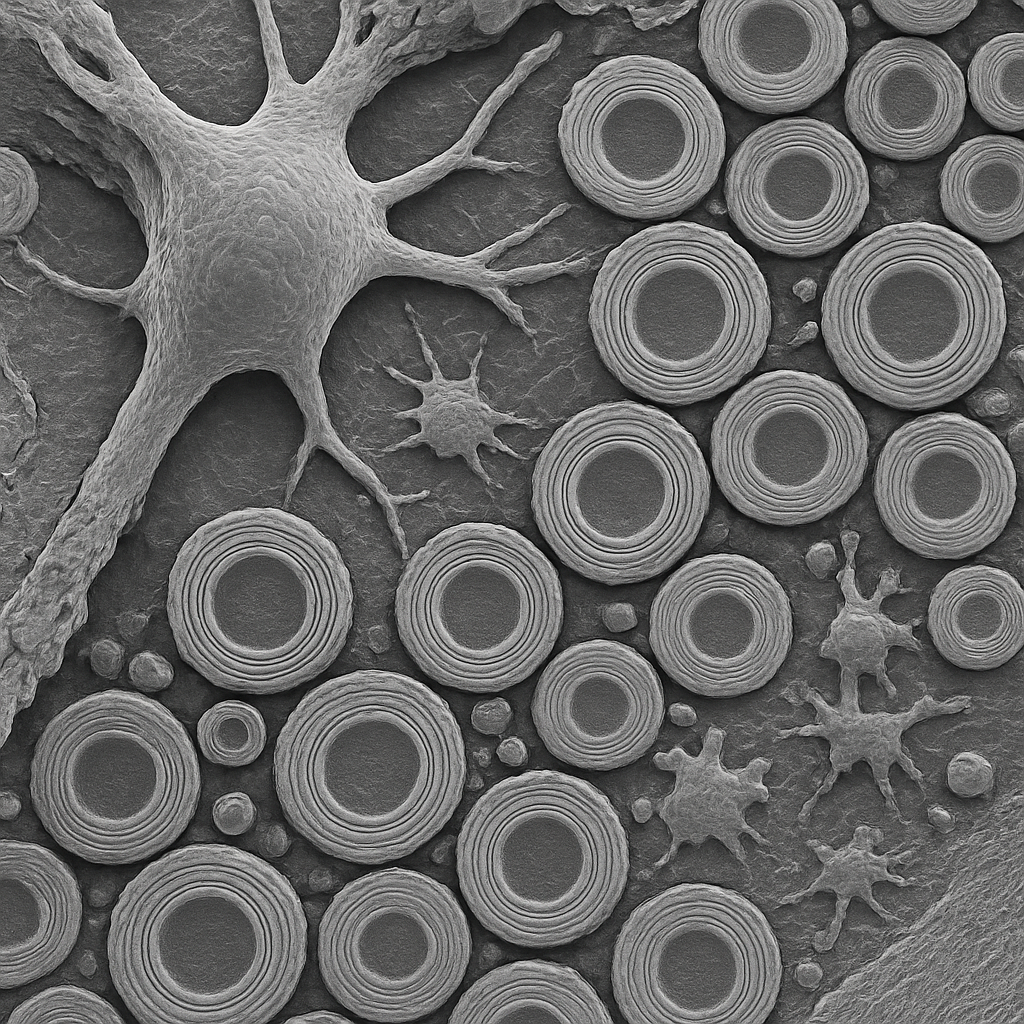
SEM at 10,000× showing a cross-fractured view through white matter revealing myelinated axon bundles in cross-section. Each axon appears as a central light core (the axoplasm) surrounded by concentric rings of dark myelin sheath. The myelin wrapping shows the characteristic lamellar structure. Axon diameters range from ~0.5 to 5 μm. Between the myelinated fibers, thin processes of oligodendrocyte cytoplasm and astrocyte processes fill the extracellular space. The freeze-fracture surface gives a sharp, textured appearance to the cut face.
10,000× FE-SEM, freeze-fractured white matter cross-section. Myelinated axons in cross-section — lamellar myelin architecture visible. Hitachi Regulus 8200 — cold cathode FE-SEM, 0.6 nm res.
OpenAI gpt-image-1
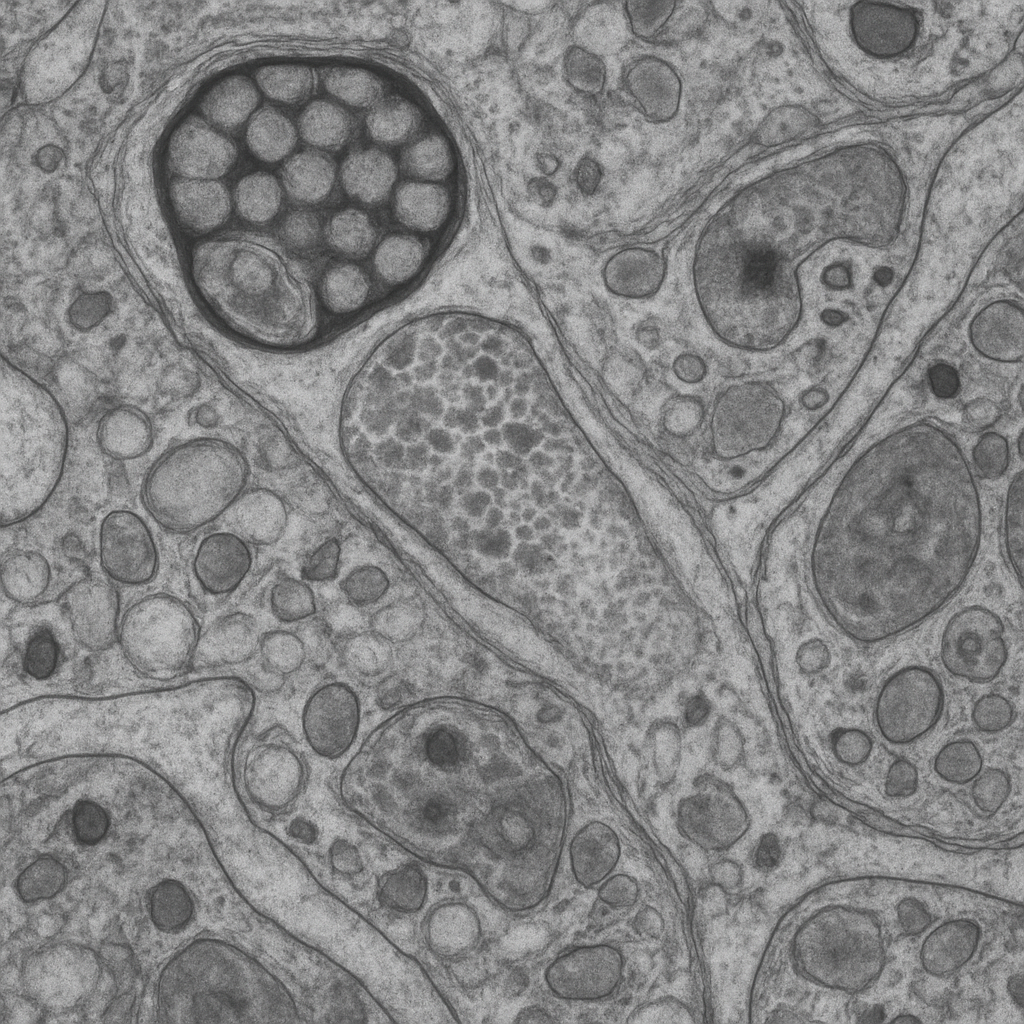
Transmission electron micrograph at 25,000× showing the ultrastructure of a chemical synapse in the cerebral cortex. The presynaptic terminal (bouton) contains a cluster of round, clear synaptic vesicles (~40 nm diameter) filled with neurotransmitter. The vesicles are concentrated at the active zone — the site of vesicle fusion and transmitter release. The synaptic cleft (~20 nm wide) separates the pre- and postsynaptic membranes. The postsynaptic membrane shows a prominent electron-dense postsynaptic density (PSD). Mitochondria with tubular cristae are present in the presynaptic terminal, providing ATP for vesicle recycling.
25,000× TEM, 200 kV. Ultrathin section (70 nm) stained with uranyl acetate and lead citrate. Synaptic vesicles, cleft, PSD, and presynaptic mitochondria all resolved. JEOL JEM-F200 — cold FEG TEM, 0.19 nm res.
OpenAI gpt-image-1
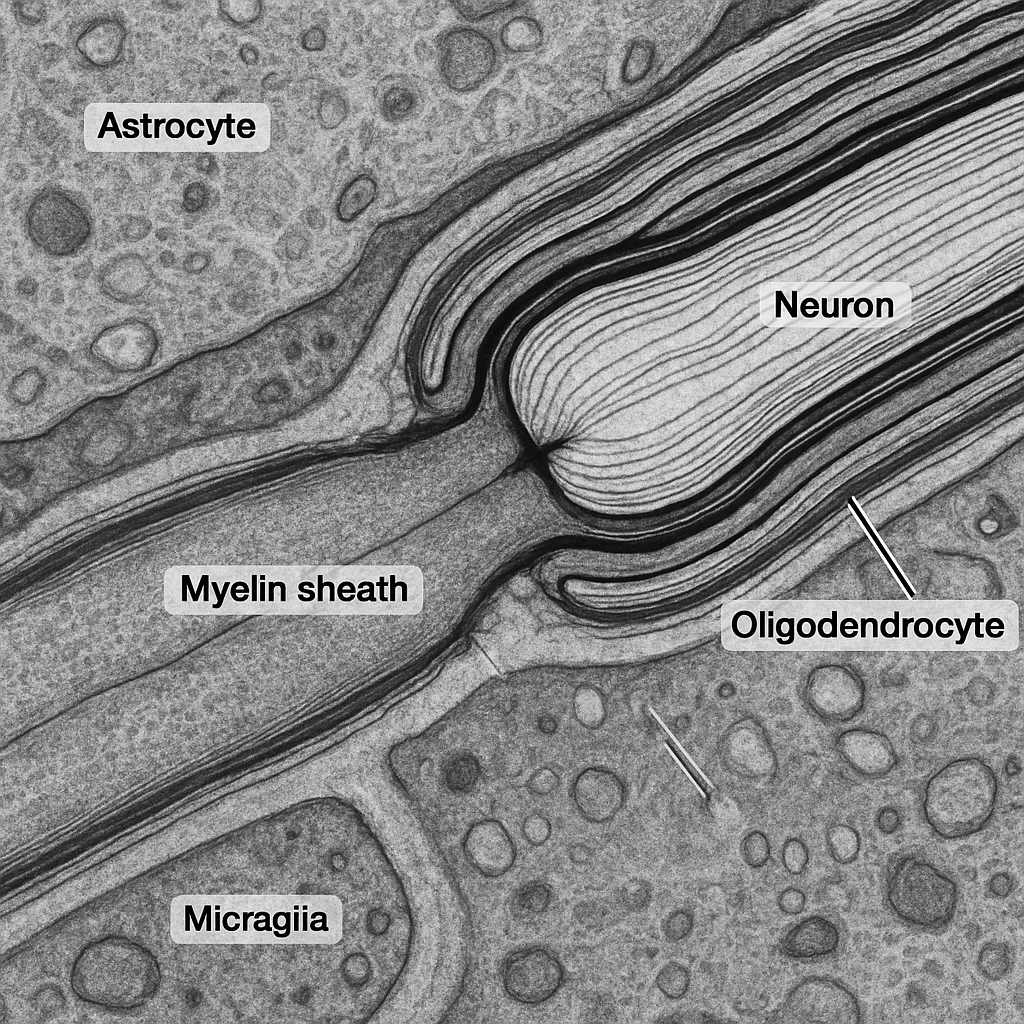
TEM at 50,000× showing a longitudinal section through a myelinated axon at a node of Ranvier. The compact myelin sheath — composed of tightly wrapped oligodendrocyte membranes — appears as alternating dark (major dense line) and light (intraperiod line) lamellae, each ~12 nm in periodicity. At the node, the myelin sheath terminates in paranodal loops that contact the axonal membrane. The bare nodal axolemma is visible as an electron-dense segment rich in voltage-gated sodium channels. The axoplasm contains neurofilaments (10 nm intermediate filaments) and microtubules (25 nm hollow cylinders) running parallel to the axon long axis.
50,000× TEM, 200 kV. Myelin lamellae with 12 nm periodicity resolved. Node of Ranvier — paranodal loops and bare nodal axolemma visible. Neurofilaments and microtubules in axoplasm. Thermo Fisher Talos F200X G2 — 0.12 nm S/TEM resolution.
OpenAI gpt-image-1
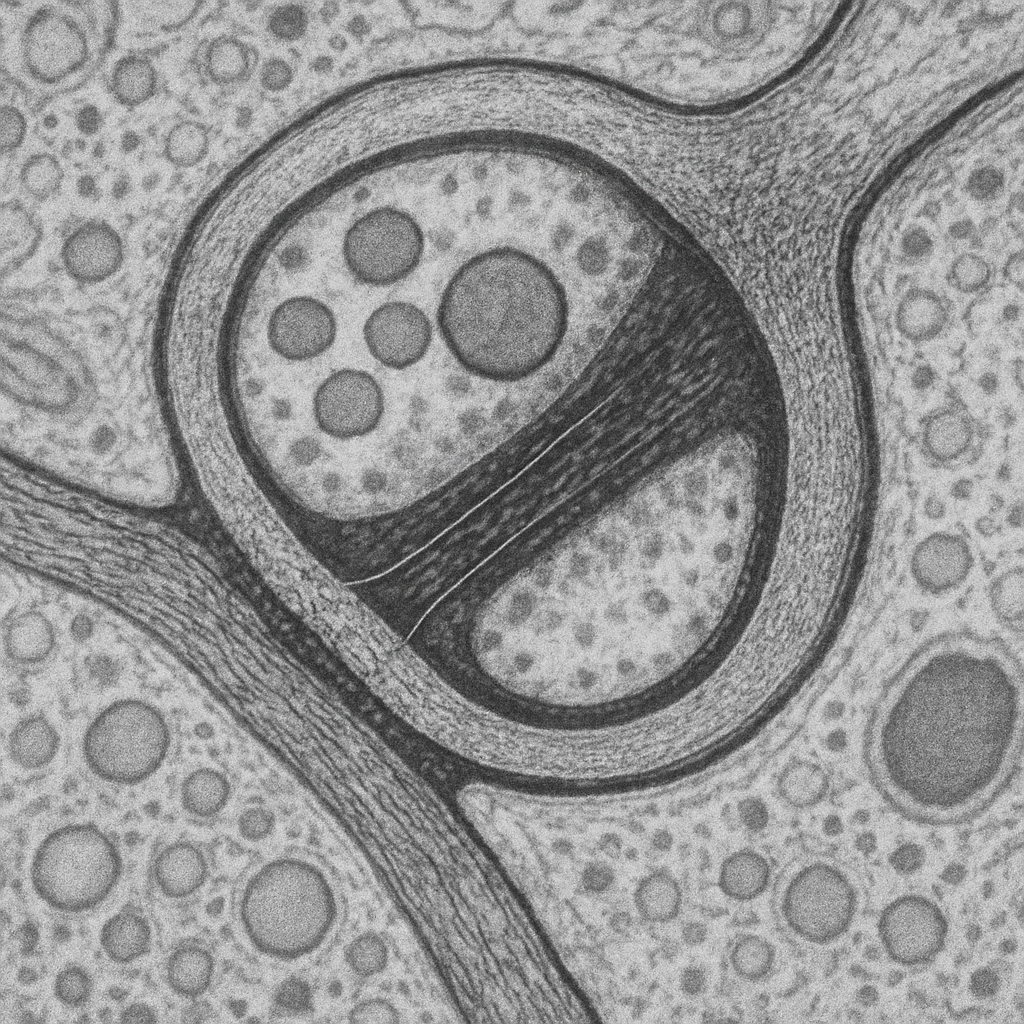
High-magnification TEM at 100,000× showing the molecular architecture of an excitatory glutamatergic synapse. Individual synaptic vesicles (~40 nm diameter) are clearly resolved as round membrane-bound structures with electron-lucent interiors. The active zone shows vesicles docked at the presynaptic membrane, with electron-dense material representing the SNARE protein complex. The synaptic cleft (~20 nm) contains filamentous material bridging the pre- and post- synaptic membranes. The postsynaptic density (PSD) is a thick electron-dense band (~30–50 nm) rich in scaffolding proteins, receptors (NMDA, AMPA), and signaling molecules. The plasma membrane lipid bilayer is resolved as a 7 nm tri-laminar structure.
100,000× TEM, 200 kV. Molecular-scale synaptic architecture — individual vesicles, SNARE docking, synaptic cleft filaments, PSD, and lipid bilayer all resolved. Thermo Fisher Talos F200X G2 — cryo-capable, 0.12 nm S/TEM res.
OpenAI gpt-image-1
Compatible Microscopes
| Model | Manufacturer | Type | Magnification Range | NA Max | Resolution |
|---|---|---|---|---|---|
| HT7800 | Hitachi High-Tech | Tem | 200–600000× | — | — nm |
| Regulus 8200 | Hitachi High-Tech | Fe Sem | 20–2000000× | — | 0.6 nm |
| JEM-F200 | JEOL | Tem | 50–1500000× | — | — nm |
| JSM-IT800 | JEOL | Sem | 5–1000000× | — | 0.7 nm |
| JSM-7900F | JEOL | Fe Sem | 25–1000000× | — | 0.5 nm |
| BZ-X810 | Keyence | Fluorescence All In One | 2–100× | 1.45 | 190 nm |
| VHX-7000 | Keyence | Digital Microscope | 0.1–6000× | — | 100 nm |
| DM6 B | Leica Microsystems | Upright Optical | 1.25–100× | 1.4 | 200 nm |
| DMi8 | Leica Microsystems | Inverted Optical | 2.5–100× | 1.47 | 185 nm |
| STELLARIS 5 | Leica Microsystems | Confocal | 5–100× | — | 120 nm |
| Eclipse Ti2 | Nikon | Inverted Optical | 2–100× | 1.45 | 190 nm |
| Eclipse Ni | Nikon | Upright Optical | 2–100× | 1.4 | 200 nm |
| A1R HD25 | Nikon | Confocal | 4–100× | — | 140 nm |
| BX53 | Olympus (Evident) | Upright Optical | 2–100× | 1.4 | 200 nm |
| IX83 | Olympus (Evident) | Inverted Optical | 2–100× | 1.4 | 200 nm |
| FV4000 | Olympus (Evident) | Confocal | 4–100× | — | 120 nm |
| Apreo 2 | Thermo Fisher Scientific | Fe Sem | 13–2000000× | — | 0.7 nm |
| Axio Observer 7 | Zeiss | Inverted Optical | 5–100× | 1.4 | 200 nm |
| Axio Imager 2 | Zeiss | Upright Optical | 1.25–100× | 1.4 | 200 nm |
| LSM 980 | Zeiss | Confocal | 5–63× | — | 120 nm |
| Primostar 3 | Zeiss | Upright Optical | 4–100× | 1.25 | 350 nm |